Development and characterization of a reverse genetic system for studying dengue virus serotype 3 strain variation and neutralization
- PMID: 22389731
- PMCID: PMC3289595
- DOI: 10.1371/journal.pntd.0001486
Development and characterization of a reverse genetic system for studying dengue virus serotype 3 strain variation and neutralization
Abstract
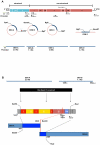
Dengue viruses (DENV) are enveloped single-stranded positive-sense RNA viruses transmitted by Aedes spp. mosquitoes. There are four genetically distinct serotypes designated DENV-1 through DENV-4, each further subdivided into distinct genotypes. The dengue scientific community has long contended that infection with one serotype confers lifelong protection against subsequent infection with the same serotype, irrespective of virus genotype. However this hypothesis is under increased scrutiny and the role of DENV genotypic variation in protection from repeated infection is less certain. As dengue vaccine trials move increasingly into field-testing, there is an urgent need to develop tools to better define the role of genotypic variation in DENV infection and immunity. To better understand genotypic variation in DENV-3 neutralization and protection, we designed and constructed a panel of isogenic, recombinant DENV-3 infectious clones, each expressing an envelope glycoprotein from a different DENV-3 genotype; Philippines 1982 (genotype I), Thailand 1995 (genotype II), Sri Lanka 1989 and Cuba 2002 (genotype III) and Puerto Rico 1977 (genotype IV). We used the panel to explore how natural envelope variation influences DENV-polyclonal serum interactions. When the recombinant viruses were tested in neutralization assays using immune sera from primary DENV infections, neutralization titers varied by as much as ∼19-fold, depending on the expressed envelope glycoprotein. The observed variability in neutralization titers suggests that relatively few residue changes in the E glycoprotein may have significant effects on DENV specific humoral immunity and influence antibody mediated protection or disease enhancement in the setting of both natural infection and vaccination. These genotypic differences are also likely to be important in temporal and spatial microevolution of DENV-3 in the background of heterotypic neutralization. The recombinant and synthetic tools described here are valuable for testing hypotheses on genetic determinants of DENV-3 immunopathogenesis.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- Kyle JL, Harris E. Global spread and persistence of dengue. Annu Rev Microbiol. 2008;62:71–92. - PubMed
-
- Burke DS, Nisalak A, Johnson DE, Scott RM. A prospective study of dengue infections in bangkok. Am J Trop Med Hyg. 1988;38:172–180. - PubMed
-
- [Anonymous] Dengue and dengue hemorrhagic fever. Wallingford, Oxon, UK; New York, , NY, USA: CAB International; 1997. Available: http://search.lib.unc.edu?R=UNCb3110999.
-
- Halstead SB. Antibodies determine virulence in dengue. Ann N Y Acad Sci. 2009;1171(Suppl 1):E48–56. - PubMed
-
- Rothman AL. Cellular immunology of sequential dengue virus infection and its role in disease pathogenesis. Curr Top Microbiol Immunol. 2010;338:83–98. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

