Regulation of global gene expression in human Loa loa infection is a function of chronicity
- PMID: 22389737
- PMCID: PMC3289604
- DOI: 10.1371/journal.pntd.0001527
Regulation of global gene expression in human Loa loa infection is a function of chronicity
Abstract
Background: Human filarial infection is characterized by downregulated parasite-antigen specific T cell responses but distinct differences exist between patients with longstanding infection (endemics) and those who acquired infection through temporary residency or visits to filarial-endemic regions (expatriates).
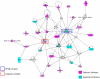
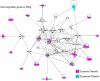
Methods and findings: To characterize mechanisms underlying differences in T cells, analysis of global gene expression using human spotted microarrays was conducted on CD4(+) and CD8(+) T cells from microfilaremic Loa loa-infected endemic and expatriate patients. Assessment of unstimulated cells showed overexpression of genes linked to inflammation and caspase-associated cell death, particularly in endemics, and enrichment of the Th1/Th2 canonical pathway in endemic CD4(+) cells. However, pathways within CD8(+) unstimulated cells were most significantly enriched in both patient groups. Antigen (Ag)-driven gene expression was assessed to microfilarial Ag (MfAg) and to the nonparasite Ag streptolysin O (SLO). For MfAg-driven cells, the number of genes differing significantly from unstimulated cells was greater in endemics compared to expatriates (p<0.0001). Functional analysis showed a differential increase in genes associated with NFkB (both groups) and caspase activation (endemics). While the expatriate response to MfAg was primarily a CD4(+) pro-inflammatory one, the endemic response included CD4(+) and CD8(+) cells and was linked to insulin signaling, histone complexes, and ubiquitination. Unlike the enrichment of canonical pathways in CD8(+) unstimulated cells, both groups showed pathway enrichment in CD4(+) cells to MfAg. Contrasting with the divergent responses to MfAg seen between endemics and expatriates, the CD4(+) response to SLO was similar; however, CD8(+) cells differed strongly in the nature and numbers (156 [endemics] vs 36 [expatriates]) of genes with differential expression.
Conclusions: These data suggest several important pathways are responsible for the different outcomes seen among filarial-infected patients with varying levels of chronicity and imply an important role for CD8(+) cells in some of the global changes seen with lifelong exposure.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Similar articles
-
Blastogenic responses, interleukin-2 production and interleukin-2 receptor expression on CD4+ and CD8+ lymphocytes in rhesus monkeys experimentally inoculated with Loa loa.Parasite Immunol. 1997 Jul;19(7):301-8. doi: 10.1046/j.1365-3024.1997.d01-212.x. Parasite Immunol. 1997. PMID: 9278942
-
Increased frequency of Th2-type cytokine-producing T cells in microfilaremic loiasis.Am J Trop Med Hyg. 1999 Apr;60(4):680-6. doi: 10.4269/ajtmh.1999.60.680. Am J Trop Med Hyg. 1999. PMID: 10348248
-
T helper responsiveness in human Loa loa infection; defective specific proliferation and cytokine production by CD4+ T cells from microfilaraemic subjects compared with amicrofilaraemics.Clin Exp Immunol. 1997 May;108(2):272-8. doi: 10.1046/j.1365-2249.1997.d01-1010.x. Clin Exp Immunol. 1997. PMID: 9158097 Free PMC article.
-
Imported Loa loa filariasis: three cases and a review of cases reported in non-endemic countries in the past 25 years.Int J Infect Dis. 2012 Sep;16(9):e649-62. doi: 10.1016/j.ijid.2012.05.1023. Epub 2012 Jul 10. Int J Infect Dis. 2012. PMID: 22784545 Review.
-
Maturation of human neonatal CD4+ and CD8+ T lymphocytes into Th1/Th2 effectors.Vaccine. 1998 Aug-Sep;16(14-15):1415-9. doi: 10.1016/s0264-410x(98)00101-7. Vaccine. 1998. PMID: 9711781 Review.
Cited by
-
The Potentials and Pitfalls of Microarrays in Neglected Tropical Diseases: A Focus on Human Filarial Infections.Microarrays (Basel). 2016 Aug 2;5(3):20. doi: 10.3390/microarrays5030020. Microarrays (Basel). 2016. PMID: 27600086 Free PMC article. Review.
-
Human innate lymphoid cells (ILCs) in filarial infections.Parasite Immunol. 2018 Feb;40(2):10.1111/pim.12442. doi: 10.1111/pim.12442. Epub 2017 Jun 15. Parasite Immunol. 2018. PMID: 28504838 Free PMC article. Review.
-
Eosinophil-associated processes underlie differences in clinical presentation of loiasis between temporary residents and those indigenous to Loa-endemic areas.Clin Infect Dis. 2015 Jan 1;60(1):55-63. doi: 10.1093/cid/ciu723. Epub 2014 Sep 18. Clin Infect Dis. 2015. PMID: 25234520 Free PMC article.
-
Highlighting the Relevance of CD8+ T Cells in Filarial Infections.Front Immunol. 2021 Sep 16;12:714052. doi: 10.3389/fimmu.2021.714052. eCollection 2021. Front Immunol. 2021. PMID: 34603287 Free PMC article. Review.
-
Looking beyond the induction of Th2 responses to explain immunomodulation by helminths.Parasite Immunol. 2015 Jun;37(6):304-13. doi: 10.1111/pim.12194. Parasite Immunol. 2015. PMID: 25869527 Free PMC article. Review.
References
-
- Klion AD, Massougbodji A, Sadeler BC, Ottesen EA, Nutman TB. Loiasis in endemic and nonendemic populations: immunologically mediated differences in clinical presentation. J Infect Dis. 1991;163:1318–1325. - PubMed
-
- McCarthy JS, Ottesen EA, Nutman TB. Onchocerciasis in endemic and nonendemic populations: differences in clinical presentation and immunologic findings. J Infect Dis. 1994;170:736–741. - PubMed
-
- Partono F, Oemijati S, Hudojo, Joesoef A, Sajidiman H, et al. Malayan filariasis in Central Sulawesi (Celebes), Indonesia. Southeast Asian J Trop Med Public Health. 1977;8:452–458. - PubMed
-
- Nutman TB, Miller KD, Mulligan M, Ottesen EA. Loa loa infection in temporary residents of endemic regions: recognition of a hyperresponsive syndrome with characteristic clinical manifestations. J Infect Dis. 1986;154:10–18. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

