Alternative splicing mediates responses of the Arabidopsis circadian clock to temperature changes
- PMID: 22408072
- PMCID: PMC3336117
- DOI: 10.1105/tpc.111.093948
Alternative splicing mediates responses of the Arabidopsis circadian clock to temperature changes
Abstract
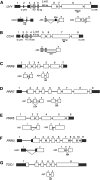
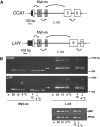
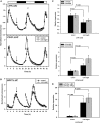
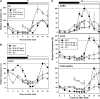
Alternative splicing plays crucial roles by influencing the diversity of the transcriptome and proteome and regulating protein structure/function and gene expression. It is widespread in plants, and alteration of the levels of splicing factors leads to a wide variety of growth and developmental phenotypes. The circadian clock is a complex piece of cellular machinery that can regulate physiology and behavior to anticipate predictable environmental changes on a revolving planet. We have performed a system-wide analysis of alternative splicing in clock components in Arabidopsis thaliana plants acclimated to different steady state temperatures or undergoing temperature transitions. This revealed extensive alternative splicing in clock genes and dynamic changes in alternatively spliced transcripts. Several of these changes, notably those affecting the circadian clock genes late elongated hypocotyl (LHY) and pseudo response regulator7, are temperature-dependent and contribute markedly to functionally important changes in clock gene expression in temperature transitions by producing nonfunctional transcripts and/or inducing nonsense-mediated decay. Temperature effects on alternative splicing contribute to a decline in LHY transcript abundance on cooling, but LHY promoter strength is not affected. We propose that temperature-associated alternative splicing is an additional mechanism involved in the operation and regulation of the plant circadian clock.
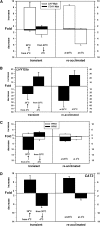
Figures




Similar articles
-
Unproductive alternative splicing and nonsense mRNAs: a widespread phenomenon among plant circadian clock genes.Biol Direct. 2012 Jul 2;7:20. doi: 10.1186/1745-6150-7-20. Biol Direct. 2012. PMID: 22747664 Free PMC article.
-
The Arabidopsis sickle Mutant Exhibits Altered Circadian Clock Responses to Cool Temperatures and Temperature-Dependent Alternative Splicing.Plant Cell. 2016 Oct;28(10):2560-2575. doi: 10.1105/tpc.16.00223. Epub 2016 Sep 13. Plant Cell. 2016. PMID: 27624757 Free PMC article.
-
A self-regulatory circuit of CIRCADIAN CLOCK-ASSOCIATED1 underlies the circadian clock regulation of temperature responses in Arabidopsis.Plant Cell. 2012 Jun;24(6):2427-42. doi: 10.1105/tpc.112.098723. Epub 2012 Jun 19. Plant Cell. 2012. PMID: 22715042 Free PMC article.
-
Regulation of the circadian clock through pre-mRNA splicing in Arabidopsis.J Exp Bot. 2014 May;65(8):1973-80. doi: 10.1093/jxb/eru085. Epub 2014 Mar 6. J Exp Bot. 2014. PMID: 24604736 Review.
-
Chromatin remodeling and alternative splicing: pre- and post-transcriptional regulation of the Arabidopsis circadian clock.Semin Cell Dev Biol. 2013 May;24(5):399-406. doi: 10.1016/j.semcdb.2013.02.009. Epub 2013 Mar 15. Semin Cell Dev Biol. 2013. PMID: 23499867 Review.
Cited by
-
Global approaches for telling time: omics and the Arabidopsis circadian clock.Semin Cell Dev Biol. 2013 May;24(5):383-92. doi: 10.1016/j.semcdb.2013.02.005. Epub 2013 Feb 20. Semin Cell Dev Biol. 2013. PMID: 23435351 Free PMC article. Review.
-
Unraveling the circadian clock in Arabidopsis.Plant Signal Behav. 2013 Feb;8(2):e23014. doi: 10.4161/psb.23014. Epub 2012 Dec 6. Plant Signal Behav. 2013. PMID: 23221775 Free PMC article.
-
Many Facets of Dynamic Plasticity in Plants.Cold Spring Harb Perspect Biol. 2019 Oct 1;11(10):a034629. doi: 10.1101/cshperspect.a034629. Cold Spring Harb Perspect Biol. 2019. PMID: 31138545 Free PMC article. Review.
-
Editorial: Molecular basis of seed germination and dormancy.Front Plant Sci. 2023 Jul 10;14:1242428. doi: 10.3389/fpls.2023.1242428. eCollection 2023. Front Plant Sci. 2023. PMID: 37492776 Free PMC article. No abstract available.
-
A sugarcane R2R3-MYB transcription factor gene is alternatively spliced during drought stress.Sci Rep. 2017 Feb 7;7:41922. doi: 10.1038/srep41922. Sci Rep. 2017. PMID: 28167824 Free PMC article.
References
-
- Alabadí D., Oyama T., Yanovsky M.J., Harmon F.G., Más P., Kay S.A. (2001). Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293: 880–883 - PubMed
-
- Arciga-Reyes L., Wootton L., Kieffer M., Davies B. (2006). UPF1 is required for nonsense-mediated mRNA decay (NMD) and RNAi in Arabidopsis. Plant J. 47: 480–489 - PubMed
-
- Barash Y., Calarco J.A., Gao W., Pan Q., Wang X., Shai O., Blencowe B.J., Frey B.J. (2010). Deciphering the splicing code. Nature 465: 53–59 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

