The structural biology of HIV-1: mechanistic and therapeutic insights
- PMID: 22421880
- PMCID: PMC3588166
- DOI: 10.1038/nrmicro2747
The structural biology of HIV-1: mechanistic and therapeutic insights
Abstract
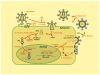
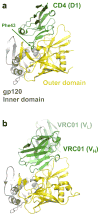
Three-dimensional molecular structures can provide detailed information on biological mechanisms and, for cases in which the molecular function affects human health, can significantly aid in the development of therapeutic interventions. For almost 25 years, key components of the lentivirus HIV-1, including the envelope glycoproteins, the capsid and the replication enzymes reverse transcriptase, integrase and protease, have been scrutinized to near atomic-scale resolution. Moreover, structural analyses of the interactions between viral and host cell components have yielded key insights into the mechanisms of viral entry, chromosomal integration, transcription and egress from cells. Here, we review recent advances in HIV-1 structural biology, focusing on the molecular mechanisms of viral replication and on the development of new therapeutics.
Figures




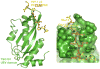
References
-
- Gao F, et al. Origin of HIV-1 in the chimpanzee Pan troglodytes troglodytes. Nature. 1999;397:436–441. - PubMed
-
- Korber B, et al. Timing the ancestor of the HIV-1 pandemic strains. Science. 2000;288:1789–1796. - PubMed
-
- Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

