Protein dynamics studied with ultrafast two-dimensional infrared vibrational echo spectroscopy
- PMID: 22433178
- PMCID: PMC3389584
- DOI: 10.1021/ar200275k
Protein dynamics studied with ultrafast two-dimensional infrared vibrational echo spectroscopy
Abstract
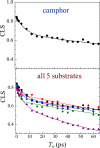
Proteins, enzymes, and other biological molecules undergo structural dynamics as an intrinsic part of their biological functions. While many biological processes occur on the millisecond, second, and even longer time scales, the fundamental structural dynamics that eventually give rise to such processes occur on much faster time scales. Many decades ago, chemical kineticists focused on the inverse of the reaction rate constant as the important time scale for a chemical reaction. However, through transition state theory and a vast amount of experimental evidence, we now know that the key events in a chemical reaction can involve structural fluctuations that take a system of reactants to its transition state, the crossing of a barrier, and the eventual relaxation to product states. Such dynamics occur on very fast time scales. Today researchers would like to investigate the fast structural fluctuations of biological molecules to gain an understanding of how biological processes proceed from simple structural changes in biomolecules to the final, complex biological function. The study of the fast structural dynamics of biological molecules requires experiments that operate on the appropriate time scales, and in this Account, we discuss the application of ultrafast two-dimensional infrared (2D IR) vibrational echo spectroscopy to the study of protein dynamics. The 2D IR vibrational echo experiment is akin to 2D NMR, but it operates on time scales many orders of magnitude faster. In the experiments, a particular vibrational oscillator serves as a vibrational dynamics probe. As the structure of the protein evolves in time, the structural changes are manifested as time-dependent changes in the frequency of the vibrational dynamics probe. The 2D IR vibrational echo experiments can track the vibrational frequency evolution, which we then relate to the time evolution of the protein structure. In particular, we measured protein substate interconversion for mutants of myoglobin using 2D IR chemical exchange spectroscopy and observed well-defined substate interconversion on a sub-100 ps time scale. In another study, we investigated the influence of binding five different substrates to the enzyme cytochrome P450(cam). The various substrates affect the enzyme dynamics differently, and the observed dynamics are correlated with the enzyme's selectivity of hydroxylation of the substrates and with the substrate binding affinity.
Figures

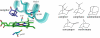

Similar articles
-
Water dynamics in salt solutions studied with ultrafast two-dimensional infrared (2D IR) vibrational echo spectroscopy.Acc Chem Res. 2009 Sep 15;42(9):1210-9. doi: 10.1021/ar900043h. Acc Chem Res. 2009. PMID: 19378969 Free PMC article.
-
Protein dynamics in cytochrome P450 molecular recognition and substrate specificity using 2D IR vibrational echo spectroscopy.J Am Chem Soc. 2011 Mar 23;133(11):3995-4004. doi: 10.1021/ja109168h. Epub 2011 Feb 24. J Am Chem Soc. 2011. PMID: 21348488 Free PMC article.
-
Two-dimensional IR spectroscopy of protein dynamics using two vibrational labels: a site-specific genetically encoded unnatural amino acid and an active site ligand.J Phys Chem B. 2011 Sep 29;115(38):11294-304. doi: 10.1021/jp206986v. Epub 2011 Aug 31. J Phys Chem B. 2011. PMID: 21823631 Free PMC article.
-
Dynamics of liquids, molecules, and proteins measured with ultrafast 2D IR vibrational echo chemical exchange spectroscopy.Annu Rev Phys Chem. 2009;60:21-38. doi: 10.1146/annurev-physchem-073108-112712. Annu Rev Phys Chem. 2009. PMID: 18851709 Review.
-
2D-IR spectroscopy: ultrafast insights into biomolecule structure and function.Chem Soc Rev. 2009 Jul;38(7):1837-48. doi: 10.1039/b819181f. Epub 2009 Mar 27. Chem Soc Rev. 2009. PMID: 19551165 Review.
Cited by
-
Molecular Vibrational Polariton Dynamics: What Can Polaritons Do?Acc Chem Res. 2023 Apr 4;56(7):776-786. doi: 10.1021/acs.accounts.2c00796. Epub 2023 Mar 17. Acc Chem Res. 2023. PMID: 36930582 Free PMC article.
-
Structural Dependence of Extended Amide III Vibrations in Two-Dimensional Infrared Spectra.J Phys Chem Lett. 2023 Oct 19;14(41):9257-9264. doi: 10.1021/acs.jpclett.3c02662. Epub 2023 Oct 9. J Phys Chem Lett. 2023. PMID: 37812580 Free PMC article.
-
Reaction dynamics studied via femtosecond X-ray liquidography at X-ray free-electron lasers.Chem Sci. 2022 Jun 6;13(29):8457-8490. doi: 10.1039/d2sc00502f. eCollection 2022 Jul 29. Chem Sci. 2022. PMID: 35974755 Free PMC article. Review.
-
Quantum process tomography quantifies coherence transfer dynamics in vibrational exciton.J Phys Chem B. 2013 Oct 31;117(43):13631-8. doi: 10.1021/jp4075493. Epub 2013 Oct 16. J Phys Chem B. 2013. PMID: 24079417 Free PMC article.
-
Exploring Conformational Landscapes Along Anharmonic Low-Frequency Vibrations.J Phys Chem B. 2024 Jul 25;128(29):7112-7120. doi: 10.1021/acs.jpcb.4c02743. Epub 2024 Jul 10. J Phys Chem B. 2024. PMID: 38986052 Free PMC article.
References
-
- Henzler-Wildman KA, Lei M, Thai V, Kerns SJ, Karplus M, Kern D. A Hierarchy of Timescales in Protein Dynamics Is Linked to Enzyme Catalysis. Nature. 2007;450:913–916. - PubMed
-
- Hammes-Schiffer S, Benkovic SJ. Relating Protein Motion to Catalysis. Annu. Rev. Biochem. 2006;75:519–541. - PubMed
-
- Erzberger JP, Berger JM. Evolutionary Relationships and Structural Mechanisms of AAA+ Proteins. Annu. Rev. Biophys. Biomol. Struct. 2006;35:93–114. - PubMed
-
- Boehr DD, Dyson HJ, Wright PE. An NMR Perspective on Enzyme Dynamics. Chem. Rev. 2006;106:3055–3079. - PubMed
-
- Frauenfelder H, Sligar SG, Wolynes PG. The Energy Landscapes and Motions of Proteins. Science. 1991;254:1598–1603. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

