Effect of image compression and scaling on automated scoring of immunohistochemical stainings and segmentation of tumor epithelium
- PMID: 22436596
- PMCID: PMC3375185
- DOI: 10.1186/1746-1596-7-29
Effect of image compression and scaling on automated scoring of immunohistochemical stainings and segmentation of tumor epithelium
Abstract
Background: Digital whole-slide scanning of tissue specimens produces large images demanding increasing storing capacity. To reduce the need of extensive data storage systems image files can be compressed and scaled down. The aim of this article is to study the effect of different levels of image compression and scaling on automated image analysis of immunohistochemical (IHC) stainings and automated tumor segmentation.
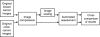
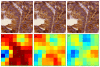
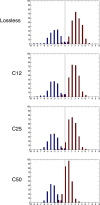
Methods: Two tissue microarray (TMA) slides containing 800 samples of breast cancer tissue immunostained against Ki-67 protein and two TMA slides containing 144 samples of colorectal cancer immunostained against EGFR were digitized with a whole-slide scanner. The TMA images were JPEG2000 wavelet compressed with four compression ratios: lossless, and 1:12, 1:25 and 1:50 lossy compression. Each of the compressed breast cancer images was furthermore scaled down either to 1:1, 1:2, 1:4, 1:8, 1:16, 1:32, 1:64 or 1:128. Breast cancer images were analyzed using an algorithm that quantitates the extent of staining in Ki-67 immunostained images, and EGFR immunostained colorectal cancer images were analyzed with an automated tumor segmentation algorithm. The automated tools were validated by comparing the results from losslessly compressed and non-scaled images with results from conventional visual assessments. Percentage agreement and kappa statistics were calculated between results from compressed and scaled images and results from lossless and non-scaled images.
Results: Both of the studied image analysis methods showed good agreement between visual and automated results. In the automated IHC quantification, an agreement of over 98% and a kappa value of over 0.96 was observed between losslessly compressed and non-scaled images and combined compression ratios up to 1:50 and scaling down to 1:8. In automated tumor segmentation, an agreement of over 97% and a kappa value of over 0.93 was observed between losslessly compressed images and compression ratios up to 1:25.
Conclusions: The results of this study suggest that images stored for assessment of the extent of immunohistochemical staining can be compressed and scaled significantly, and images of tumors to be segmented can be compressed without compromising computer-assisted analysis results using studied methods.
Virtual slides: The virtual slide(s) for this article can be found here: http://www.diagnosticpathology.diagnomx.eu/vs/2442925476534995.
Figures


References
-
- Konsti J, Lundin J, Jumppanen M, Lundin M, Viitanen A, Isola J. A public-domain image processing tool for automated quantification of fluorescence in situ hybridisation signals. J Clin Pathol. 2008;61:278–282. - PubMed
-
- Konsti J, Lundin M, Joensuu H, Lehtimaki T, Sihto H, Holli K, Turpeenniemi-Hujanen T, Kataja V, Sailas L, Isola J, Lundin J. Development and evaluation of a virtual microscopy application for automated assessment of Ki-67 expression in breast cancer. BMC Clin Pathol. 2011;11:3. doi: 10.1186/1472-6890-11-3. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

