H2A.Z.2.2 is an alternatively spliced histone H2A.Z variant that causes severe nucleosome destabilization
- PMID: 22467210
- PMCID: PMC3401452
- DOI: 10.1093/nar/gks267
H2A.Z.2.2 is an alternatively spliced histone H2A.Z variant that causes severe nucleosome destabilization
Abstract
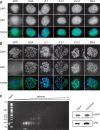
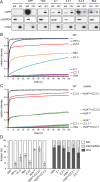
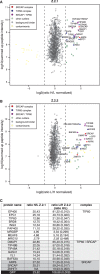
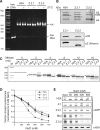
The histone variant H2A.Z has been implicated in many biological processes, such as gene regulation and genome stability. Here, we present the identification of H2A.Z.2.2 (Z.2.2), a novel alternatively spliced variant of histone H2A.Z and provide a comprehensive characterization of its expression and chromatin incorporation properties. Z.2.2 mRNA is found in all human cell lines and tissues with highest levels in brain. We show the proper splicing and in vivo existence of this variant protein in humans. Furthermore, we demonstrate the binding of Z.2.2 to H2A.Z-specific TIP60 and SRCAP chaperone complexes and its active replication-independent deposition into chromatin. Strikingly, various independent in vivo and in vitro analyses, such as biochemical fractionation, comparative FRAP studies of GFP-tagged H2A variants, size exclusion chromatography and single molecule FRET, in combination with in silico molecular dynamics simulations, consistently demonstrate that Z.2.2 causes major structural changes and significantly destabilizes nucleosomes. Analyses of deletion mutants and chimeric proteins pinpoint this property to its unique C-terminus. Our findings enrich the list of known human variants by an unusual protein belonging to the H2A.Z family that leads to the least stable nucleosome known to date.
Figures




References
-
- van Holde KE. Chromatin. New York: Springer; 1988.
-
- Bonisch C, Nieratschker SM, Orfanos NK, Hake SB. Chromatin proteomics and epigenetic regulatory circuits. Exp. Rev. Proteomics. 2008;5:105–119. - PubMed
-
- Bernstein E, Hake SB. The nucleosome: a little variation goes a long way. Biochem. Cell Biol. 2006;84:505–517. - PubMed
-
- Boulard M, Bouvet P, Kundu TK, Dimitrov S. Histone variant nucleosomes: structure, function and implication in disease. Subcell. Biochem. 2007;41:71–89. - PubMed
-
- Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W. Histone H2A variants H2AX and H2AZ. Curr. Opin. Genet. Dev. 2002;12:162–169. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

