Confirmation of an epilepsy modifier locus on mouse chromosome 11 and candidate gene analysis by RNA-Seq
- PMID: 22471526
- PMCID: PMC3370141
- DOI: 10.1111/j.1601-183X.2012.00790.x
Confirmation of an epilepsy modifier locus on mouse chromosome 11 and candidate gene analysis by RNA-Seq
Abstract
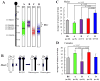
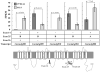
Epilepsy is a neurological disorder affecting approximately 1% of the worldwide population. Mutations in voltage-gated sodium channels have been identified in several monogenic epilepsy syndromes. Over 800 mutations have been identified in the voltage-gated sodium channel genes SCN1A and SCN2A in human epilepsies, including genetic epilepsy with febrile seizures plus (GEFS+) and Dravet syndrome. In GEFS+ families, affected members with the same mutation often display variability in clinical severity of the disease. This suggests that additional genes modify the effect of the primary mutation, resulting in the variable clinical presentation. The Scn2a(Q54) transgenic mouse model has an epilepsy phenotype that varies depending on the genetic strain background. Scn2a(Q54) mice congenic on the C57BL/6J strain exhibit delayed seizure onset and improved survival compared to (C57BL/6J × SJL/J)F1.Q54 mice. Two modifier loci of Scn2a(Q54) seizure susceptibility were mapped and designated Moe1 (modifier of epilepsy) on chromosome (chr) 11 and Moe2 on chr 19. To confirm Moe1 and refine its position, we generated interval-specific congenic lines carrying C57BL/6J-derived chr 11 alleles on the SJL/J strain and refined the map position to 89-104 Mb. We then used RNA-Seq for candidate analysis in the modifier region. C57BL/6J and SJL/J male and female brain RNAs were sequenced, revealing numerous significant transcriptome differences and coding single-nucleotide polymorphisms. Additional consideration of gene function and expression suggested several strong candidate modifier genes, including two voltage-gated calcium channel subunits, Cacna1g and Cacnb1, and the proline and acidic amino acid-rich basic leucine zipper transcription factor, Hlf.
© 2012 The Authors. Genes, Brain and Behavior © 2012 Blackwell Publishing Ltd and International Behavioural and Neural Genetics Society.
Figures


References
-
- Beck H, Steffens R, Elger CE, Heinemann U. Voltage-dependent Ca2+ currents in epilepsy. Epilepsy Res. 1998;32:321–332. - PubMed
-
- Bergren SK, Chen S, Galecki A, Kearney JA. Genetic modifiers affecting severity of epilepsy caused by mutation of sodium channel Scn2a. Mamm Genome. 2005;16:683–690. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

