Formal modeling and analysis of the MAL-associated biological regulatory network: insight into cerebral malaria
- PMID: 22479409
- PMCID: PMC3316585
- DOI: 10.1371/journal.pone.0033532
Formal modeling and analysis of the MAL-associated biological regulatory network: insight into cerebral malaria
Abstract
The discrete modeling formalism of René Thomas is a well known approach for the modeling and analysis of Biological Regulatory Networks (BRNs). This formalism uses a set of parameters which reflect the dynamics of the BRN under study. These parameters are initially unknown but may be deduced from the appropriately chosen observed dynamics of a BRN. The discrete model can be further enriched by using the model checking tool HyTech along with delay parameters. This paves the way to accurately analyse a BRN and to make predictions about critical trajectories which lead to a normal or diseased response. In this paper, we apply the formal discrete and hybrid (discrete and continuous) modeling approaches to characterize behavior of the BRN associated with MyD88-adapter-like (MAL)--a key protein involved with innate immune response to infections. In order to demonstrate the practical effectiveness of our current work, different trajectories and corresponding conditions that may lead to the development of cerebral malaria (CM) are identified. Our results suggest that the system converges towards hyperinflammation if Bruton's tyrosine kinase (BTK) remains constitutively active along with pre-existing high cytokine levels which may play an important role in CM pathogenesis.
Conflict of interest statement
Figures

 B as I
B as I B is degraded. The proinflammatory cytokine genes are expressed (6) producing INCY that are secreted (7). INCY are responsible (8) for the production of inflammation and activation of their respective receptors. This again activates NF-
B is degraded. The proinflammatory cytokine genes are expressed (6) producing INCY that are secreted (7). INCY are responsible (8) for the production of inflammation and activation of their respective receptors. This again activates NF- B (9a) and through an alternate pathway induces the production of SOCS-1 (9b). SOCS-1 negatively regulates MAL by polyubiquitination (10a) and blocks NF-
B (9a) and through an alternate pathway induces the production of SOCS-1 (9b). SOCS-1 negatively regulates MAL by polyubiquitination (10a) and blocks NF- B mediated expression (10b). Abbreviations: TLR, toll like receptors; PAMPs, pathogen associated molecular patterns; BTK, bruton's tyrosine kinase; MAL, MyD88 adapter like; MyD88, myeloid differentiation response protein; NF-
B mediated expression (10b). Abbreviations: TLR, toll like receptors; PAMPs, pathogen associated molecular patterns; BTK, bruton's tyrosine kinase; MAL, MyD88 adapter like; MyD88, myeloid differentiation response protein; NF- B, nuclear factor kappa-light-chain-enhancer of activated B cells; I
B, nuclear factor kappa-light-chain-enhancer of activated B cells; I B, inhibitor of
B, inhibitor of  B; INCY, proinflammatory cytokines; SOCS-1, suppressor of cytokine signaling-1 , , .
B; INCY, proinflammatory cytokines; SOCS-1, suppressor of cytokine signaling-1 , , .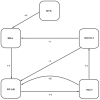

 ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  ,
,  and
and  .
.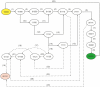
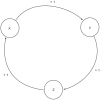
 ,
,  and
and  represent biological entities. The labels ‘+’ and ‘1’ represent the activation and the threshold concentration respectively.
represent biological entities. The labels ‘+’ and ‘1’ represent the activation and the threshold concentration respectively.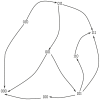
 ,
,  and
and  .
.
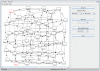


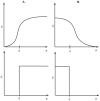
 is shown as: sigmoidal representation of the activation (A.), discrete approximation of the activation (B.), piece-wise linear approximation of the activation (C.), sigmoidal representation of the degradation (D.), discrete approximation of the degradation (E.) and piece-wise linear approximation of the degradation (F.).
is shown as: sigmoidal representation of the activation (A.), discrete approximation of the activation (B.), piece-wise linear approximation of the activation (C.), sigmoidal representation of the degradation (D.), discrete approximation of the degradation (E.) and piece-wise linear approximation of the degradation (F.).
References
-
- Who malaria fact sheet no. 94 world health organization. URL http://www.who.int/mediacentre/factsheets/fs094/en/index.html. Accessed April 2010.
-
- Miller LH, Baruch DI, Marsh K, Doumbo OK. The pathogenic basis of malaria. Nature. 2002;415:673–679. - PubMed
-
- Clark IA, Cowden WB. Why is the pathology of falciparum worse than that of vivax malaria? Parasitol Today (Regul Ed) 1999;15:458–461. - PubMed
-
- Amante FH, Haque A, Stanley AC, Rivera FdeL, Randall LM, et al. Immune-mediated mechanisms of parasite tissue sequestration during experimental cerebral malaria. J Immunol. 2010;185:3632–3642. - PubMed
-
- Hunt NH, Golenser J, Chan-Ling T, Parekh S, Rae C, et al. Immunopathogenesis of cerebral malaria. Int J Parasitol. 2006;36:569–582. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

