Asymmetric mode of Ca²⁺-S100A4 interaction with nonmuscle myosin IIA generates nanomolar affinity required for filament remodeling
- PMID: 22483112
- PMCID: PMC3343272
- DOI: 10.1016/j.str.2012.02.002
Asymmetric mode of Ca²⁺-S100A4 interaction with nonmuscle myosin IIA generates nanomolar affinity required for filament remodeling
Abstract
Filament assembly of nonmuscle myosin IIA (NMIIA) is selectively regulated by the small Ca²⁺-binding protein, S100A4, which causes enhanced cell migration and metastasis in certain cancers. Our NMR structure shows that an S100A4 dimer binds to a single myosin heavy chain in an asymmetrical configuration. NMIIA in the complex forms a continuous helix that stretches across the surface of S100A4 and engages the Ca²⁺-dependent binding sites of each subunit in the dimer. Synergy between these sites leads to a very tight association (K(D) ∼1 nM) that is unique in the S100 family. Single-residue mutations that remove this synergy weaken binding and ameliorate the effects of S100A4 on NMIIA filament assembly and cell spreading in A431 human epithelial carcinoma cells. We propose a model for NMIIA filament disassembly by S100A4 in which initial binding to the unstructured NMIIA tail initiates unzipping of the coiled coil and disruption of filament packing.
Copyright © 2012 Elsevier Ltd. All rights reserved.
Figures

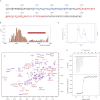

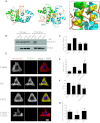


References
-
- Ambartsumian N.S., Grigorian M.S., Larsen I.F., Karlstrøm O., Sidenius N., Rygaard J., Georgiev G., Lukanidin E. Metastasis of mammary carcinomas in GRS/A hybrid mice transgenic for the mts1 gene. Oncogene. 1996;13:1621–1630. - PubMed
-
- Atkinson S.J., Stewart M. Molecular interactions in myosin assembly. Role of the 28-residue charge repeat in the rod. J. Mol. Biol. 1992;226:7–13. - PubMed
-
- Bhattacharya S., Large E., Heizmann C.W., Hemmings B.A., Chazin W.J. Structure of the Ca2+/S100B/NDR kinase peptide complex: insights into S100 target specificity and activation of the kinase. Biochemistry. 2003;42:14416–14426. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

