Related bifunctional restriction endonuclease-methyltransferase triplets: TspDTI, Tth111II/TthHB27I and TsoI with distinct specificities
- PMID: 22489904
- PMCID: PMC3384240
- DOI: 10.1186/1471-2199-13-13
Related bifunctional restriction endonuclease-methyltransferase triplets: TspDTI, Tth111II/TthHB27I and TsoI with distinct specificities
Abstract
Background: We previously defined a family of restriction endonucleases (REases) from Thermus sp., which share common biochemical and biophysical features, such as the fusion of both the nuclease and methyltransferase (MTase) activities in a single polypeptide, cleavage at a distance from the recognition site, large molecular size, modulation of activity by S-adenosylmethionine (SAM), and incomplete cleavage of the substrate DNA. Members include related thermophilic REases with five distinct specificities: TspGWI, TaqII, Tth111II/TthHB27I, TspDTI and TsoI.
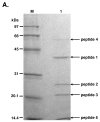
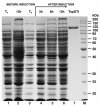
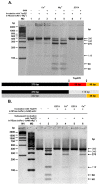
Results: TspDTI, TsoI and isoschizomers Tth111II/TthHB27I recognize different, but related sequences: 5'-ATGAA-3', 5'-TARCCA-3' and 5'-CAARCA-3' respectively. Their amino acid sequences are similar, which is unusual among REases of different specificity. To gain insight into this group of REases, TspDTI, the prototype member of the Thermus sp. enzyme family, was cloned and characterized using a recently developed method for partially cleaving REases.
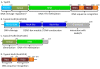
Conclusions: TspDTI, TsoI and isoschizomers Tth111II/TthHB27I are closely related bifunctional enzymes. They comprise a tandem arrangement of Type I-like domains, like other Type IIC enzymes (those with a fusion of a REase and MTase domains), e.g. TspGWI, TaqII and MmeI, but their sequences are only remotely similar to these previously characterized enzymes. The characterization of TspDTI, a prototype member of this group, extends our understanding of sequence-function relationships among multifunctional restriction-modification enzymes.
Figures


Similar articles
-
Three-stage biochemical selection: cloning of prototype class IIS/IIC/IIG restriction endonuclease-methyltransferase TsoI from the thermophile Thermus scotoductus.BMC Mol Biol. 2013 Aug 6;14:17. doi: 10.1186/1471-2199-14-17. BMC Mol Biol. 2013. PMID: 23919831 Free PMC article.
-
A new prototype IIS/IIC/IIG endonuclease-methyltransferase TsoI from the thermophile Thermus scotoductus, recognising 5'-TARCCA(N11/9)-3' sequences.J Biotechnol. 2015 Jan 20;194:19-26. doi: 10.1016/j.jbiotec.2014.11.023. Epub 2014 Dec 4. J Biotechnol. 2015. PMID: 25481098
-
Two-stage gene assembly/cloning of a member of the TspDTI subfamily of bifunctional restriction endonucleases, TthHB27I.J Biotechnol. 2015 Jan 20;194:67-80. doi: 10.1016/j.jbiotec.2014.11.030. Epub 2014 Dec 6. J Biotechnol. 2015. PMID: 25486633
-
Restriction endonucleases: classification, properties, and applications.Mol Biotechnol. 2003 Mar;23(3):225-43. doi: 10.1385/mb:23:3:225. Mol Biotechnol. 2003. PMID: 12665693 Review.
-
Type II restriction endonucleases: structure and mechanism.Cell Mol Life Sci. 2005 Mar;62(6):685-707. doi: 10.1007/s00018-004-4513-1. Cell Mol Life Sci. 2005. PMID: 15770420 Free PMC article. Review.
Cited by
-
A new genomic tool, ultra-frequently cleaving TaqII/sinefungin endonuclease with a combined 2.9-bp recognition site, applied to the construction of horse DNA libraries.BMC Genomics. 2013 Jun 1;14:370. doi: 10.1186/1471-2164-14-370. BMC Genomics. 2013. PMID: 23724933 Free PMC article.
-
Engineering TaqII bifunctional endonuclease DNA recognition fidelity: the effect of a single amino acid substitution within the methyltransferase catalytic site.Mol Biol Rep. 2016 Apr;43(4):269-82. doi: 10.1007/s11033-016-3949-3. Epub 2016 Feb 17. Mol Biol Rep. 2016. PMID: 26886214
-
Thermostable proteins bioprocesses: The activity of restriction endonuclease-methyltransferase from Thermus thermophilus (RM.TthHB27I) cloned in Escherichia coli is critically affected by the codon composition of the synthetic gene.PLoS One. 2017 Oct 17;12(10):e0186633. doi: 10.1371/journal.pone.0186633. eCollection 2017. PLoS One. 2017. PMID: 29040308 Free PMC article.
-
Three-stage biochemical selection: cloning of prototype class IIS/IIC/IIG restriction endonuclease-methyltransferase TsoI from the thermophile Thermus scotoductus.BMC Mol Biol. 2013 Aug 6;14:17. doi: 10.1186/1471-2199-14-17. BMC Mol Biol. 2013. PMID: 23919831 Free PMC article.
-
Cofactor analogue-induced chemical reactivation of endonuclease activity in a DNA cleavage/methylation deficient TspGWI N₄₇₃A variant in the NPPY motif.Mol Biol Rep. 2014;41(4):2313-23. doi: 10.1007/s11033-014-3085-x. Epub 2014 Jan 19. Mol Biol Rep. 2014. PMID: 24442320 Free PMC article.
References
-
- Szybalski W, Kim SC, Hasan N, Podhajska AJ. Class-IIS restriction enzymes - a review. Gene. 1991;100:13–26. - PubMed
-
- The Restriction Enzyme Database. http://rebase.neb.com
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

