MSV3d: database of human MisSense Variants mapped to 3D protein structure
- PMID: 22491796
- PMCID: PMC3317913
- DOI: 10.1093/database/bas018
MSV3d: database of human MisSense Variants mapped to 3D protein structure
Abstract
The elucidation of the complex relationships linking genotypic and phenotypic variations to protein structure is a major challenge in the post-genomic era. We present MSV3d (Database of human MisSense Variants mapped to 3D protein structure), a new database that contains detailed annotation of missense variants of all human proteins (20 199 proteins). The multi-level characterization includes details of the physico-chemical changes induced by amino acid modification, as well as information related to the conservation of the mutated residue and its position relative to functional features in the available or predicted 3D model. Major releases of the database are automatically generated and updated regularly in line with the dbSNP (database of Single Nucleotide Polymorphism) and SwissVar releases, by exploiting the extensive Décrypthon computational grid resources. The database (http://decrypthon.igbmc.fr/msv3d) is easily accessible through a simple web interface coupled to a powerful query engine and a standard web service. The content is completely or partially downloadable in XML or flat file formats. Database URL: http://decrypthon.igbmc.fr/msv3d.
Figures

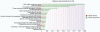

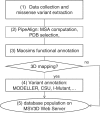
References
-
- Chasman D, Adams RM. Predicting the functional consequences of non-synonymous single nucleotide polymorphisms: structure-based assessment of amino acid variation. J. Mol. Biol. 2001;307:683–706. - PubMed
-
- Thusberg J, Vihinen M. Pathogenic or not? And if so, then how? Studying the effects of missense mutations using bioinformatics methods. Hum. Mutat. 2009;30:703–714. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

