Direct ubiquitin independent recognition and degradation of a folded protein by the eukaryotic proteasomes-origin of intrinsic degradation signals
- PMID: 22506054
- PMCID: PMC3323579
- DOI: 10.1371/journal.pone.0034864
Direct ubiquitin independent recognition and degradation of a folded protein by the eukaryotic proteasomes-origin of intrinsic degradation signals
Abstract
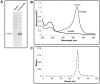
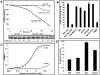
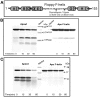
Eukaryotic 26S proteasomes are structurally organized to recognize, unfold and degrade globular proteins. However, all existing model substrates of the 26S proteasome in addition to ubiquitin or adaptor proteins require unstructured regions in the form of fusion tags for efficient degradation. We report for the first time that purified 26S proteasome can directly recognize and degrade apomyoglobin, a globular protein, in the absence of ubiquitin, extrinsic degradation tags or adaptor proteins. Despite a high affinity interaction, absence of a ligand and presence of only helices/loops that follow the degradation signal, apomyoglobin is degraded slowly by the proteasome. A short floppy F-helix exposed upon ligand removal and in conformational equilibrium with a disordered structure is mandatory for recognition and initiation of degradation. Holomyoglobin, in which the helix is buried, is neither recognized nor degraded. Exposure of the floppy F-helix seems to sensitize the proteasome and primes the substrate for degradation. Using peptide panning and competition experiments we speculate that initial encounters through the floppy helix and additional strong interactions with N-terminal helices anchors apomyoglobin to the proteasome. Stabilizing helical structure in the floppy F-helix slows down degradation. Destabilization of adjacent helices accelerates degradation. Unfolding seems to follow the mechanism of helix unraveling rather than global unfolding. Our findings while confirming the requirement for unstructured regions in degradation offers the following new insights: a) origin and identification of an intrinsic degradation signal in the substrate, b) identification of sequences in the native substrate that are likely to be responsible for direct interactions with the proteasome, and c) identification of critical rate limiting steps like exposure of the intrinsic degron and destabilization of an unfolding intermediate that are presumably catalyzed by the ATPases. Apomyoglobin emerges as a new model substrate to further explore the role of ATPases and protein structure in proteasomal degradation.
Conflict of interest statement
Figures




Similar articles
-
Ubiquitin-Like Proteasome System Represents a Eukaryotic-Like Pathway for Targeted Proteolysis in Archaea.mBio. 2016 May 17;7(3):e00379-16. doi: 10.1128/mBio.00379-16. mBio. 2016. PMID: 27190215 Free PMC article.
-
Ubiquitin recognition by the proteasome.J Biochem. 2017 Feb 1;161(2):113-124. doi: 10.1093/jb/mvw091. J Biochem. 2017. PMID: 28069863 Review.
-
The Cdc48 unfoldase prepares well-folded protein substrates for degradation by the 26S proteasome.Commun Biol. 2019 Jan 21;2:29. doi: 10.1038/s42003-019-0283-z. eCollection 2019. Commun Biol. 2019. PMID: 30675527 Free PMC article.
-
The unfolding of substrates and ubiquitin-independent protein degradation by proteasomes.Biochimie. 2001 Mar-Apr;83(3-4):311-8. doi: 10.1016/s0300-9084(01)01244-5. Biochimie. 2001. PMID: 11295491 Review.
-
Structural Insights into Substrate Recognition and Processing by the 20S Proteasome.Biomolecules. 2021 Jan 24;11(2):148. doi: 10.3390/biom11020148. Biomolecules. 2021. PMID: 33498876 Free PMC article. Review.
Cited by
-
Control of Pim2 kinase stability and expression in transformed human haematopoietic cells.Biosci Rep. 2015 Oct 23;35(6):e00274. doi: 10.1042/BSR20150217. Biosci Rep. 2015. PMID: 26500282 Free PMC article.
-
Snapshots of urea-induced early structural changes and unfolding of an ankyrin repeat protein at atomic resolution.Protein Sci. 2022 Dec;31(12):e4515. doi: 10.1002/pro.4515. Protein Sci. 2022. PMID: 36382986 Free PMC article.
-
Glucose restriction in Saccharomyces cerevisiae modulates the phosphorylation pattern of the 20S proteasome and increases its activity.Sci Rep. 2023 Nov 8;13(1):19383. doi: 10.1038/s41598-023-46614-x. Sci Rep. 2023. PMID: 37938622 Free PMC article.
-
Design principles that protect the proteasome from self-destruction.Protein Sci. 2022 Mar;31(3):556-567. doi: 10.1002/pro.4251. Epub 2021 Dec 16. Protein Sci. 2022. PMID: 34878680 Free PMC article.
-
N-Terminal Coiled-Coil Structure of ATPase Subunits of 26S Proteasome Is Crucial for Proteasome Function.PLoS One. 2015 Jul 24;10(7):e0134056. doi: 10.1371/journal.pone.0134056. eCollection 2015. PLoS One. 2015. PMID: 26208326 Free PMC article.
References
-
- Glickman MH, Ciechanover A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev. 2002;82:373–428. - PubMed
-
- Wolf DH, Hilt W. The proteasome: a proteolytic nanomachine of cell regulation and waste disposal. Biochim Biophys Acta. 2004;1695:19–31. - PubMed
-
- Kisselev AF, Akopian TN, Woo KM, Goldberg AL. The sizes of peptides generated from protein by mammalian 26 and 20 S proteasomes. Implications for understanding the degradative mechanism and antigen presentation. J Biol Chem. 1999;274:3363–3371. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

