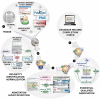
Text mining for the biocuration workflow
- PMID: 22513129
- PMCID: PMC3328793
- DOI: 10.1093/database/bas020
Text mining for the biocuration workflow
Abstract
Molecular biology has become heavily dependent on biological knowledge encoded in expert curated biological databases. As the volume of biological literature increases, biocurators need help in keeping up with the literature; (semi-) automated aids for biocuration would seem to be an ideal application for natural language processing and text mining. However, to date, there have been few documented successes for improving biocuration throughput using text mining. Our initial investigations took place for the workshop on 'Text Mining for the BioCuration Workflow' at the third International Biocuration Conference (Berlin, 2009). We interviewed biocurators to obtain workflows from eight biological databases. This initial study revealed high-level commonalities, including (i) selection of documents for curation; (ii) indexing of documents with biologically relevant entities (e.g. genes); and (iii) detailed curation of specific relations (e.g. interactions); however, the detailed workflows also showed many variabilities. Following the workshop, we conducted a survey of biocurators. The survey identified biocurator priorities, including the handling of full text indexed with biological entities and support for the identification and prioritization of documents for curation. It also indicated that two-thirds of the biocuration teams had experimented with text mining and almost half were using text mining at that time. Analysis of our interviews and survey provide a set of requirements for the integration of text mining into the biocuration workflow. These can guide the identification of common needs across curated databases and encourage joint experimentation involving biocurators, text mining developers and the larger biomedical research community.
Figures
References
-
- Lanzen A, Oinn T. The Taverna Interaction Service: enabling manual interaction in workflows. Bioinformatics. 2008;24:1118–1120. - PubMed
-
- Burns GAPC, Krallinger M, Cohen KB, et al. Biocuration Workflow Catalogue—Text Mining for the Biocuration Workflow. 2009. Nature Precedings. http://dx.doi.org/10.1038/npre.2009.3250.1 (2 March 2012, date last accessed) - DOI
-
- Krallinger M. A Framework for BioCuration Workflows (part II) Nature Precedings. 2009 http://dx.doi.org/10.1038/npre.2009.3126.1 (2 March 2012, date last accessed) - DOI
Publication types
MeSH terms
Grants and funding
- 1G08LM10720-01/LM/NLM NIH HHS/United States
- 1R01RR024031/RR/NCRR NIH HHS/United States
- R01ES014065-04S1/ES/NIEHS NIH HHS/United States
- P20RR016463/RR/NCRR NIH HHS/United States
- R01 GM083871/GM/NIGMS NIH HHS/United States
- G08 LM010720/LM/NLM NIH HHS/United States
- BB/F010486/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- R01-GM083871/GM/NIGMS NIH HHS/United States
- 2U01HG02712-04/HG/NHGRI NIH HHS/United States
- HG001315/HG/NHGRI NIH HHS/United States
- R01ES014065/ES/NIEHS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials