Comparison of DNA extraction methods for microbial community profiling with an application to pediatric bronchoalveolar lavage samples
- PMID: 22514642
- PMCID: PMC3326054
- DOI: 10.1371/journal.pone.0034605
Comparison of DNA extraction methods for microbial community profiling with an application to pediatric bronchoalveolar lavage samples
Abstract
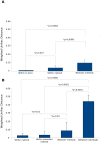
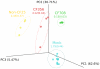
Barcoded amplicon sequencing is rapidly becoming a standard method for profiling microbial communities, including the human respiratory microbiome. While this approach has less bias than standard cultivation, several steps can introduce variation including the type of DNA extraction method used. Here we assessed five different extraction methods on pediatric bronchoalveolar lavage (BAL) samples and a mock community comprised of nine bacterial genera to determine method reproducibility and detection limits for these typically low complexity communities. Additionally, using the mock community, we were able to evaluate contamination and select a relative abundance cut-off threshold based on the geometric distribution that optimizes the trade off between detecting bona fide operational taxonomic units and filtering out spurious ones. Using this threshold, the majority of genera in the mock community were predictably detected by all extraction methods including the hard-to-lyse Gram-positive genus Staphylococcus. Differences between extraction methods were significantly greater than between technical replicates for both the mock community and BAL samples emphasizing the importance of using a standardized methodology for microbiome studies. However, regardless of method used, individual patients retained unique diagnostic profiles. Furthermore, despite being stored as raw frozen samples for over five years, community profiles from BAL samples were consistent with historical culturing results. The culture-independent profiling of these samples also identified a number of anaerobic genera that are gaining acceptance as being part of the respiratory microbiome. This study should help guide researchers to formulate sampling, extraction and analysis strategies for respiratory and other human microbiome samples.
Conflict of interest statement
Figures



References
-
- Tringe SG, Hugenholtz P. A renaissance for the pioneering 16 S rRNA gene. Curr Opin Microbiol. 2008;11:442–446. doi: 10.1016/j.mib.2008.09.011. - DOI - PubMed
-
- Guss AM, Roeselers G, Newton ILG, Young CR, Klepac-Ceraj V, et al. Phylogenetic and metabolic diversity of bacteria associated with cystic fibrosis. ISME J. 2011;5:20–29. doi: 10.1038/ismej.2010.88. - DOI - PMC - PubMed
-
- Erb-Downward JR, Thompson DL, Han MK, Freeman CM, McCloskey L, et al. Analysis of the Lung Microbiome in the “Healthy” Smoker and in COPD. PLoS ONE. 2011;6:e16384. doi: 10.1371/journal.pone.0016384. - DOI - PMC - PubMed
-
- Hilty M, Burke C, Pedro H, Cardenas P, Bush A, et al. Disordered Microbial Communities in Asthmatic Airways. PLoS ONE. 2010;5:e8578. doi: 10.1371/journal.pone.0008578. - DOI - PMC - PubMed
-
- Willner D, Haynes MR, Furlan M, Schmieder R, Lim YW, et al. Spatial distribution of microbial communities in the cystic fibrosis lung. ISME J. 2011 Available: http://dx.doi.org/10.1038/ismej.2011.104. Accessed 17 Dec 2011. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

