Transient aggregation of ubiquitinated proteins is a cytosolic unfolded protein response to inflammation and endoplasmic reticulum stress
- PMID: 22518844
- PMCID: PMC3366003
- DOI: 10.1074/jbc.M112.350934
Transient aggregation of ubiquitinated proteins is a cytosolic unfolded protein response to inflammation and endoplasmic reticulum stress
Abstract
Failure to maintain protein homeostasis (proteostasis) leads to accumulation of unfolded proteins and contributes to the pathogenesis of many human diseases. Accumulation of unfolded proteins in the endoplasmic reticulum (ER) elicits unfolded protein response (UPR) that serves to attenuate protein translation, and increase protein refolding or degradation. In contrast to UPR in the ER, the regulatory molecules operative in cytosolic responses and their potential relation to ER stress are not well elucidated. Aggresome-like induced structures (ALIS) have been described as transient aggregation of ubiquitinated proteins in the cytosol. In this study, we show that cells respond to inflammation, infection or ER stress by cytosolic formation of ALIS, indicating that ALIS formation represents an early event in cellular adjustment to altered proteostasis that occurs under these conditions. This response was aided by rapid transcriptional up-regulation of polyubiqutin-binding protein p62. NF-κB and mTOR activation were also required for ALIS formation. Importantly, we show a cross talk between UPR in the ER and cytosolic ALIS. Down-regulation of ER UPR in XBP1 deficient cells increases cyotosolic ALIS formation. Furthermore, lysosomal activity but not macroautophagy is responsible for ALIS clearance. This study reveals the underlying regulatory mechanisms of ALIS formation and clearance, and provides a previously unrecognized common adaptive mechanism for cellular responses against inflammation and ER stress.
Figures
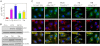

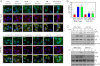
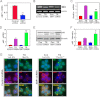

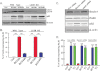

References
-
- Buchberger A., Bukau B., Sommer T. (2010) Protein quality control in the cytosol and the endoplasmic reticulum: brothers in arms. Mol. Cell 40, 238–252 - PubMed
-
- Ron D., Walter P. (2007) Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 8, 519–529 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

