Extended analysis of a genome-wide association study in primary sclerosing cholangitis detects multiple novel risk loci
- PMID: 22521342
- PMCID: PMC3399030
- DOI: 10.1016/j.jhep.2012.03.031
Extended analysis of a genome-wide association study in primary sclerosing cholangitis detects multiple novel risk loci
Abstract
Background & aims: A limited number of genetic risk factors have been reported in primary sclerosing cholangitis (PSC). To discover further genetic susceptibility factors for PSC, we followed up on a second tier of single nucleotide polymorphisms (SNPs) from a genome-wide association study (GWAS).
Methods: We analyzed 45 SNPs in 1221 PSC cases and 3508 controls. The association results from the replication analysis and the original GWAS (715 PSC cases and 2962 controls) were combined in a meta-analysis comprising 1936 PSC cases and 6470 controls. We performed an analysis of bile microbial community composition in 39 PSC patients by 16S rRNA sequencing.
Results: Seventeen SNPs representing 12 distinct genetic loci achieved nominal significance (p(replication) <0.05) in the replication. The most robust novel association was detected at chromosome 1p36 (rs3748816; p(combined)=2.1 × 10(-8)) where the MMEL1 and TNFRSF14 genes represent potential disease genes. Eight additional novel loci showed suggestive evidence of association (p(repl) <0.05). FUT2 at chromosome 19q13 (rs602662; p(comb)=1.9 × 10(-6), rs281377; p(comb)=2.1 × 10(-6) and rs601338; p(comb)=2.7 × 10(-6)) is notable due to its implication in altered susceptibility to infectious agents. We found that FUT2 secretor status and genotype defined by rs601338 significantly influence biliary microbial community composition in PSC patients.
Conclusions: We identify multiple new PSC risk loci by extended analysis of a PSC GWAS. FUT2 genotype needs to be taken into account when assessing the influence of microbiota on biliary pathology in PSC.
Copyright © 2012 European Association for the Study of the Liver. Published by Elsevier B.V. All rights reserved.
Conflict of interest statement
Figures
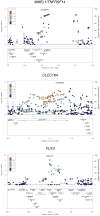


References
-
- Karlsen TH, Schrumpf E, Boberg KM. Update on primary sclerosing cholangitis. Dig Liver Dis. 2010;42:390–400. - PubMed
-
- Saarinen S, Olerup O, Broome U. Increased frequency of autoimmune diseases in patients with primary sclerosing cholangitis. Am J Gastroenterol. 2000;95:3195–3199. - PubMed
-
- Karlsen TH, Franke A, Melum E, Kaser A, Hov JR, Balschun T, et al. Genome-wide association analysis in primary sclerosing cholangitis. Gastroenterology. 2010;138:1102–1111. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

