Agouti revisited: transcript quantification of the ASIP gene in bovine tissues related to protein expression and localization
- PMID: 22530003
- PMCID: PMC3328439
- DOI: 10.1371/journal.pone.0035282
Agouti revisited: transcript quantification of the ASIP gene in bovine tissues related to protein expression and localization
Abstract
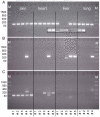
Beside its role in melanogenesis, the agouti signaling protein (ASIP) has been related to obesity. The potentially crucial role in adipocyte development makes it a tempting candidate for economic relevant, fat related traits in farm animals. The objective of our study was to characterize the mRNA expression of different ASIP transcripts and of putative targets in different bovine tissues, as well as to study consequences on protein abundance and localization. ASIP mRNA abundance was determined by RT-qPCR in adipose and further tissues of cattle representing different breeds and crosses. ASIP mRNA was up-regulated more than 9-fold in intramuscular fat of Japanese Black cattle compared to Holstein (p<0.001). Further analyses revealed that a transposon-derived transcript was solely responsible for the increased ASIP mRNA abundance. This transcript was observed in single individuals of different breeds indicating a wide spread occurrence of this insertion at the ASIP locus in cattle. The protein was detected in different adipose tissues, skin, lung and liver, but not in skeletal muscle by Western blot with a bovine-specific ASIP antibody. However, the protein abundance was not related to the observed ASIP mRNA over-expression. Immuno-histochemical analyses revealed a putative nuclear localization of ASIP additionally to the expected cytosolic signal in different cell types. The expression of melanocortin receptors (MCR) 1 to 5 as potential targets for ASIP was analyzed by RT-PCR in subcutaneous fat. Only MC1R and MC4R were detected indicating a similar receptor expression like in human adipose tissue. Our results provide evidence for a widespread expression of ASIP in bovine tissues at mRNA and, for the first time, at protein level. ASIP protein is detectable in adipocytes as well as in further cells of adipose tissue. We generated a basis for a more detailed investigation of ASIP function in peripheral tissues of various mammalian species.
Conflict of interest statement
Figures


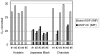



References
-
- Claycombe KJ, Xue BZ, Mynatt RL, Zemel MB, Moustaid-Moussa N. Regulation of leptin by agouti. Physiol Genomics. 2000;2:101–105. - PubMed
-
- Lu D, Willard D, Patel IR, Kadwell S, Overton L, et al. Agouti protein is an antagonist of the melanocyte-stimulating-hormone receptor. Nature. 1994;371:799–802. - PubMed
-
- Cone RD, Lu D, Koppula S, Vage DI, Klungland H, et al. The melanocortin receptors: agonists, antagonists, and the hormonal control of pigmentation. Recent Prog Horm Res. 1996;51:287–317. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

