Clonal deletion and the fate of autoreactive thymocytes that survive negative selection
- PMID: 22544394
- PMCID: PMC3362677
- DOI: 10.1038/ni.2292
Clonal deletion and the fate of autoreactive thymocytes that survive negative selection
Abstract
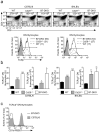
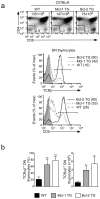
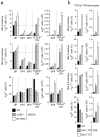
Clonal deletion of autoreactive thymocytes is important for self-tolerance, but the intrathymic signals that induce clonal deletion have not been clearly identified. We now report that clonal deletion during negative selection required CD28-mediated costimulation of autoreactive thymocytes at the CD4(+)CD8(lo) intermediate stage of differentiation. Autoreactive thymocytes were prevented from undergoing clonal deletion by either a lack of CD28 costimulation or transgenic overexpression of the antiapoptotic factors Bcl-2 or Mcl-1, with surviving thymocytes differentiating into anergic CD4(-)CD8(-) double-negative thymocytes positive for the T cell antigen receptor αβ subtype (TCRαβ) that 'preferentially' migrated to the intestine, where they re-expressed CD8α and were sequestered as CD8αα(+) intraepithelial lymphocytes (IELs). Our study identifies costimulation by CD28 as the intrathymic signal required for clonal deletion and identifies CD8αα(+) IELs as the developmental fate of autoreactive thymocytes that survive negative selection.
Figures





References
-
- Starr TK, Jameson SC, Hogquist KA. Positive and negative selection of T cells. Annual review of immunology. 2003;21:139–176. - PubMed
-
- McCaughtry TM, Hogquist KA. Central tolerance: what have we learned from mice? Semin Immunopathol. 2008;30:399–409. - PubMed
-
- Kappler JW, Roehm N, Marrack P. T cell tolerance by clonal elimination in the thymus. Cell. 1987;49:273–280. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

