Fine-scale population recombination rates, hotspots, and correlates of recombination in the Medicago truncatula genome
- PMID: 22554552
- PMCID: PMC3381680
- DOI: 10.1093/gbe/evs046
Fine-scale population recombination rates, hotspots, and correlates of recombination in the Medicago truncatula genome
Abstract
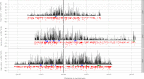
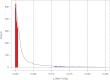
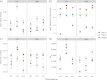
Recombination rates vary across the genome and in many species show significant relationships with several genomic features, including distance to the centromere, gene density, and GC content. Studies of fine-scale recombination rates have also revealed that in several species, there are recombination hotspots, that is, short regions with recombination rates 10-100 greater than those in surrounding regions. In this study, we analyzed whole-genome resequence data from 26 accessions of the model legume Medicago truncatula to gain insight into the genomic features that are related to high- and low-recombination rates and recombination hotspots at 1 kb scales. We found that high-recombination regions (1-kb windows among those in the highest 5% of the distribution) on all three chromosomes were significantly closer to the centromere, had higher gene density, and lower GC content than low-recombination windows. High-recombination windows are also significantly overrepresented among some gene functional categories-most strongly NB-ARC and LRR genes, both of which are important in plant defense against pathogens. Similar to high-recombination windows, recombination hotspots (1-kb windows with significantly higher recombination than the surrounding region) are significantly nearer to the centromere than nonhotspot windows. By contrast, we detected no difference in gene density or GC content between hotspot and nonhotspot windows. Using linear model wavelet analysis to examine the relationship between recombination and genomic features across multiple spatial scales, we find a significant negative correlation with distance to the centromere across scales up to 512 kb, whereas gene density and GC content show significantly positive and negative correlations, respectively, only up to 64 kb. Correlations between recombination and genomic features, particularly gene density and polymorphism, suggest that they are scale dependent and need to be assessed at scales relevant to the evolution of those features.
Figures

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

