Chromatin signatures of active enhancers
- PMID: 22555596
- PMCID: PMC3383566
- DOI: 10.4161/nucl.19232
Chromatin signatures of active enhancers
Abstract
Gene-distal cis-regulatory sequences, such as enhancers, are key contributors of tissue-specific gene expression. In particular, enhancers can be located up to hundreds of kilobases from the promoters that they control, making their identification challenging. Thanks to the recent technological advances to map histone modifications and chromatin-associated factors genome-wide, several studies have begun to characterize chromatin signatures of active enhancers. Here, we discuss some of these results and how they provide new insights into the tissue-specific organization of enhancer repertoires.
Figures
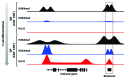
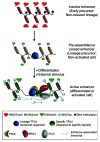
Comment on
- Pekowska A, Benoukraf T, Zacarias-Cabeza J, Belhocine M, Koch F, Holota H, et al. H3K4 tri-methylation provides an epigenetic signature of active enhancers. EMBO J. 2011;30:4198–210. doi: 10.1038/emboj.2011.295.
Similar articles
-
Evaluating Enhancer Function and Transcription.Annu Rev Biochem. 2020 Jun 20;89:213-234. doi: 10.1146/annurev-biochem-011420-095916. Epub 2020 Mar 20. Annu Rev Biochem. 2020. PMID: 32197056 Review.
-
A unique chromatin signature uncovers early developmental enhancers in humans.Nature. 2011 Feb 10;470(7333):279-83. doi: 10.1038/nature09692. Epub 2010 Dec 15. Nature. 2011. PMID: 21160473 Free PMC article.
-
The chromatin signatures of enhancers and their dynamic regulation.Nucleus. 2023 Dec;14(1):2160551. doi: 10.1080/19491034.2022.2160551. Nucleus. 2023. PMID: 36602897 Free PMC article. Review.
-
A Comprehensive Toolbox to Analyze Enhancer-Promoter Functions.Methods Mol Biol. 2021;2351:3-22. doi: 10.1007/978-1-0716-1597-3_1. Methods Mol Biol. 2021. PMID: 34382181 Review.
-
The histone variant H2A.Z is an important regulator of enhancer activity.Nucleic Acids Res. 2015 Nov 16;43(20):9742-56. doi: 10.1093/nar/gkv825. Epub 2015 Aug 28. Nucleic Acids Res. 2015. PMID: 26319018 Free PMC article.
Cited by
-
The Transcription Factor Tbx5-Dependent Epigenetic Modification Contributes to Neuropathic Allodynia by Activating TRPV1 Expression in the Dorsal Horn.J Neurosci. 2024 Sep 25;44(39):e0497242024. doi: 10.1523/JNEUROSCI.0497-24.2024. J Neurosci. 2024. PMID: 39174351 Free PMC article.
-
The basic helix-loop-helix transcription factor SHARP1 is an oncogenic driver in MLL-AF6 acute myelogenous leukemia.Nat Commun. 2018 Apr 24;9(1):1622. doi: 10.1038/s41467-018-03854-0. Nat Commun. 2018. PMID: 29692408 Free PMC article.
-
Preclinical evidence for the therapeutic value of TBX5 normalization in arrhythmia control.Cardiovasc Res. 2021 Jul 7;117(8):1908-1922. doi: 10.1093/cvr/cvaa239. Cardiovasc Res. 2021. PMID: 32777030 Free PMC article.
-
Transcription Factor Hepatocyte Nuclear Factor-1β Regulates Renal Cholesterol Metabolism.J Am Soc Nephrol. 2016 Aug;27(8):2408-21. doi: 10.1681/ASN.2015060607. Epub 2015 Dec 28. J Am Soc Nephrol. 2016. PMID: 26712526 Free PMC article.
-
Enhancers regulate genes linked to severe and mild childhood asthma.Heliyon. 2024 Jul 9;10(14):e34386. doi: 10.1016/j.heliyon.2024.e34386. eCollection 2024 Jul 30. Heliyon. 2024. PMID: 39108895 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
