Identification and characterization of the lamprey high-mobility group box 1 gene
- PMID: 22563397
- PMCID: PMC3338530
- DOI: 10.1371/journal.pone.0035755
Identification and characterization of the lamprey high-mobility group box 1 gene
Abstract
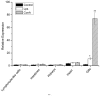
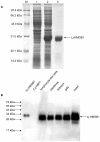
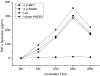
High-mobility group box 1 (HMGB1), a highly conserved DNA-binding protein, plays an important role in maintaining nucleosome structures, transcription, and inflammation. We identified a homolog of HMGB1 in the Japanese lamprey (Lampetra japonica). The Lampetra japonica HMGB1 gene (Lj-HMGB1) has over 70% sequence identity with its homologs in jawed vertebrates. Despite the reasonably high sequence identity with other HMGB1 proteins, Lj-HMGB1 did not group together with these proteins in a phylogenetic analysis. We examined Lj-HMGB1 expression in lymphocyte-like cells, and the kidneys, heart, gills, and intestines of lampreys before and after the animals were challenged with lipopolysaccharide (LPS) and concanavalin A (ConA). Lj-HMGB1 was initially expressed at a higher level in the heart, but after treatment with LPS and ConA only the gills demonstrated a significant up-regulation of expression. The recombinant Lj-HMGB1 (rLj-HMGB1) protein bound double-stranded DNA and induced the proliferation of human adenocarcinoma cells to a similar extent as human HMGB1. We further revealed that Lj-HMGB1 was able to induce the production of tumor necrosis factor-α (TNF-α), a pro-inflammatory mediator, in activated human acute monocytic leukemia cells. These results suggest that lampreys use HMGB1 to activate their innate immunity for the purpose of pathogen defense.
Conflict of interest statement
Figures
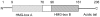





References
-
- Thomas JO, Travers AA. HMG1 and 2, and related ‘architectural’ DNA-binding proteins. Trends Biochem Sci. 2001;26:167–174. - PubMed
-
- Stott K, Tang GS, Lee KB, Thomas JO. Structure of a complex of tandem HMG boxes and DNA. J Mol Biol. 2006;360:90–104. - PubMed
-
- Topalova D, Ugrinova I, Pashev IG, Pasheva EA. HMGB1 protein inhibits DNA replication in vitro: a role of the acetylation and the acidic tail. Int J Biochem Cell Biol. 2008;40:1536–1542. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

