The human c1q globular domain: structure and recognition of non-immune self ligands
- PMID: 22566881
- PMCID: PMC3342031
- DOI: 10.3389/fimmu.2011.00092
The human c1q globular domain: structure and recognition of non-immune self ligands
Abstract
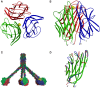
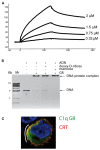
C1q, the ligand-binding unit of the C1 complex of complement, is a pattern recognition molecule with the unique ability to sense an amazing variety of targets, including a number of altered structures from self, such as apoptotic cells. The three-dimensional structure of its C-terminal globular domain, responsible for its recognition function, has been solved by X-ray crystallography, revealing a tightly packed heterotrimeric assembly with marked differences in the surface patterns of the subunits, and yielding insights into its versatile binding properties. In conjunction with other approaches, this same technique has been used recently to decipher the mechanisms that allow this domain to interact with various non-immune self ligands, including molecules known to provide eat-me signals on apoptotic cells, such as phosphatidylserine and DNA. These investigations provide evidence for a common binding area for these ligands located in subunit C of the C1q globular domain, and suggest that ligand recognition through this area down-regulates C1 activation, hence contributing to the control of the inflammatory reaction. The purpose of this article is to give an overview of these advances which represent a first step toward understanding the recognition mechanisms of C1q and their biological implications.
Keywords: C1q; X-ray crystallography; complement; innate immunity; ligand recognition.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources

