RI-1: a chemical inhibitor of RAD51 that disrupts homologous recombination in human cells
- PMID: 22573178
- PMCID: PMC3424541
- DOI: 10.1093/nar/gks353
RI-1: a chemical inhibitor of RAD51 that disrupts homologous recombination in human cells
Abstract
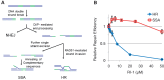
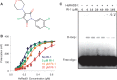
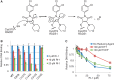
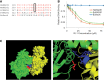
Homologous recombination serves multiple roles in DNA repair that are essential for maintaining genomic stability. We here describe RI-1, a small molecule that inhibits the central recombination protein RAD51. RI-1 specifically reduces gene conversion in human cells while stimulating single strand annealing. RI-1 binds covalently to the surface of RAD51 protein at cysteine 319 that likely destabilizes an interface used by RAD51 monomers to oligomerize into filaments on DNA. Correspondingly, the molecule inhibits the formation of subnuclear RAD51 foci in cells following DNA damage, while leaving replication protein A focus formation unaffected. Finally, it potentiates the lethal effects of a DNA cross-linking drug in human cells. Given that this inhibitory activity is seen in multiple human tumor cell lines, RI-1 holds promise as an oncologic drug. Furthermore, RI-1 represents a unique tool to dissect the network of reaction pathways that contribute to DNA repair in cells.
Figures
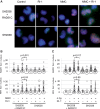

Similar articles
-
An optimized RAD51 inhibitor that disrupts homologous recombination without requiring Michael acceptor reactivity.J Med Chem. 2013 Jan 10;56(1):254-63. doi: 10.1021/jm301565b. Epub 2012 Dec 26. J Med Chem. 2013. PMID: 23231413 Free PMC article.
-
New RAD51 Inhibitors to Target Homologous Recombination in Human Cells.Genes (Basel). 2021 Jun 16;12(6):920. doi: 10.3390/genes12060920. Genes (Basel). 2021. PMID: 34208492 Free PMC article.
-
The homologous recombination protein RAD51 is a promising therapeutic target for cervical carcinoma.Oncol Rep. 2017 Aug;38(2):767-774. doi: 10.3892/or.2017.5724. Epub 2017 Jun 15. Oncol Rep. 2017. PMID: 28627709 Free PMC article.
-
Inhibiting homologous recombination by targeting RAD51 protein.Biochim Biophys Acta Rev Cancer. 2021 Dec;1876(2):188597. doi: 10.1016/j.bbcan.2021.188597. Epub 2021 Jul 28. Biochim Biophys Acta Rev Cancer. 2021. PMID: 34332021 Review.
-
Novel insights into RAD51 activity and regulation during homologous recombination and DNA replication.Biochem Cell Biol. 2016 Oct;94(5):407-418. doi: 10.1139/bcb-2016-0012. Epub 2016 Mar 31. Biochem Cell Biol. 2016. PMID: 27224545 Free PMC article. Review.
Cited by
-
Direct evidence of CRISPR-Cas9-mediated mitochondrial genome editing.Innovation (Camb). 2022 Sep 27;3(6):100329. doi: 10.1016/j.xinn.2022.100329. eCollection 2022 Nov 8. Innovation (Camb). 2022. PMID: 36275864 Free PMC article.
-
A high-throughput chemical screen with FDA approved drugs reveals that the antihypertensive drug Spironolactone impairs cancer cell survival by inhibiting homology directed repair.Nucleic Acids Res. 2014 May;42(9):5689-701. doi: 10.1093/nar/gku217. Epub 2014 Mar 25. Nucleic Acids Res. 2014. PMID: 24682826 Free PMC article.
-
The trans cell cycle effects of PARP inhibitors underlie their selectivity toward BRCA1/2-deficient cells.Genes Dev. 2021 Sep 1;35(17-18):1271-1289. doi: 10.1101/gad.348479.121. Epub 2021 Aug 12. Genes Dev. 2021. PMID: 34385259 Free PMC article.
-
Tax impairs DNA replication forks and increases DNA breaks in specific oncogenic genome regions.Mol Cancer. 2014 Sep 4;13:205. doi: 10.1186/1476-4598-13-205. Mol Cancer. 2014. PMID: 25185513 Free PMC article.
-
Identification and Characterization of Novel Compounds Blocking Shiga Toxin Expression in Escherichia coli O157:H7.Front Microbiol. 2016 Nov 30;7:1930. doi: 10.3389/fmicb.2016.01930. eCollection 2016. Front Microbiol. 2016. PMID: 27965652 Free PMC article.
References
-
- Liu N, Lamerdin JE, Tebbs RS, Schild D, Tucker JD, Shen MR, Brookman KW, Siciliano MJ, Walter CA, Fan W, et al. XRCC2 and XRCC3, new human Rad51-family members, promote chromosome stability and protect against DNA cross-links and other damages. Mol. Cell. 1998;1:783–793. - PubMed
-
- Thompson LH, Schild D. Homologous recombinational repair of DNA ensures mammalian chromosome stability. Mutat. Res. 2001;477:131–153. - PubMed
-
- Martin RW, Orelli BJ, Yamazoe M, Minn AJ, Takeda S, Bishop DK. RAD51 up-regulation bypasses BRCA1 function and is a common feature of BRCA1-deficient breast tumors. Cancer Res. 2007;67:9658–9665. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

