RTEL1 contributes to DNA replication and repair and telomere maintenance
- PMID: 22593209
- PMCID: PMC3395665
- DOI: 10.1091/mbc.E12-03-0179
RTEL1 contributes to DNA replication and repair and telomere maintenance
Abstract
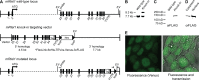
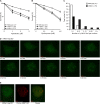
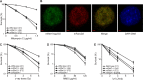
Telomere maintenance and DNA repair are important processes that protect the genome against instability. mRtel1, an essential helicase, is a dominant factor setting telomere length in mice. In addition, mRtel1 is involved in DNA double-strand break repair. The role of mRtel1 in telomere maintenance and genome stability is poorly understood. Therefore we used mRtel1-deficient mouse embryonic stem cells to examine the function of mRtel1 in replication, DNA repair, recombination, and telomere maintenance. mRtel1-deficient mouse embryonic stem cells showed sensitivity to a range of DNA-damaging agents, highlighting its role in replication and genome maintenance. Deletion of mRtel1 increased the frequency of sister chromatid exchange events and suppressed gene replacement, demonstrating the involvement of the protein in homologous recombination. mRtel1 localized transiently at telomeres and is needed for efficient telomere replication. Of interest, in the absence of mRtel1, telomeres in embryonic stem cells appeared relatively stable in length, suggesting that mRtel1 is required to allow extension by telomerase. We propose that mRtel1 is a key protein for DNA replication, recombination, and repair and efficient elongation of telomeres by telomerase.
Figures
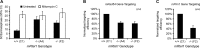
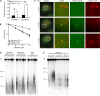
Similar articles
-
Human RTEL1 deficiency causes Hoyeraal-Hreidarsson syndrome with short telomeres and genome instability.Hum Mol Genet. 2013 Aug 15;22(16):3239-49. doi: 10.1093/hmg/ddt178. Epub 2013 Apr 15. Hum Mol Genet. 2013. PMID: 23591994
-
The mINO80 chromatin remodeling complex is required for efficient telomere replication and maintenance of genome stability.Cell Res. 2013 Dec;23(12):1396-413. doi: 10.1038/cr.2013.113. Epub 2013 Aug 27. Cell Res. 2013. PMID: 23979016 Free PMC article.
-
Stabilization of Reversed Replication Forks by Telomerase Drives Telomere Catastrophe.Cell. 2018 Jan 25;172(3):439-453.e14. doi: 10.1016/j.cell.2017.11.047. Epub 2017 Dec 28. Cell. 2018. PMID: 29290468 Free PMC article.
-
Resolving Roadblocks to Telomere Replication.Methods Mol Biol. 2019;1999:31-57. doi: 10.1007/978-1-4939-9500-4_2. Methods Mol Biol. 2019. PMID: 31127568 Review.
-
Consequences of telomere replication failure: the other end-replication problem.Trends Biochem Sci. 2022 Jun;47(6):506-517. doi: 10.1016/j.tibs.2022.03.013. Epub 2022 Apr 16. Trends Biochem Sci. 2022. PMID: 35440402 Free PMC article. Review.
Cited by
-
Human regulator of telomere elongation helicase 1 (RTEL1) is required for the nuclear and cytoplasmic trafficking of pre-U2 RNA.Nucleic Acids Res. 2015 Feb 18;43(3):1834-47. doi: 10.1093/nar/gku1402. Epub 2015 Jan 27. Nucleic Acids Res. 2015. PMID: 25628358 Free PMC article.
-
Beyond Telomerase: Telomere Instability as a Novel Target for Cancer Therapy.J Mol Genet Med. 2013 Dec;7(4):91. doi: 10.4172/1747-0862.1000091. Epub 2013 Dec 9. J Mol Genet Med. 2013. PMID: 27123041 Free PMC article.
-
Analysis of difference of association between polymorphisms in the XRCC5, RPA3 and RTEL1 genes and glioma, astrocytoma and glioblastoma.Am J Cancer Res. 2015 Jun 15;5(7):2294-300. eCollection 2015. Am J Cancer Res. 2015. PMID: 26328260 Free PMC article.
-
SMARCAL1 Resolves Replication Stress at ALT Telomeres.Cell Rep. 2016 Feb 9;14(5):1032-1040. doi: 10.1016/j.celrep.2016.01.011. Epub 2016 Jan 28. Cell Rep. 2016. PMID: 26832416 Free PMC article.
-
Solving the Telomere Replication Problem.Genes (Basel). 2017 Jan 31;8(2):55. doi: 10.3390/genes8020055. Genes (Basel). 2017. PMID: 28146113 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

