Phages and HIV-1: from display to interplay
- PMID: 22606007
- PMCID: PMC3344243
- DOI: 10.3390/ijms13044727
Phages and HIV-1: from display to interplay
Abstract
The complex hide-and-seek game between HIV-1 and the host immune system has impaired the development of an efficient vaccine. In addition, the high variability of the virus impedes the long-term control of viral replication by small antiviral drugs. For more than 20 years, phage display technology has been intensively used in the field of HIV-1 to explore the epitope landscape recognized by monoclonal and polyclonal HIV-1-specific antibodies, thereby providing precious data about immunodominant and neutralizing epitopes. In parallel, biopanning experiments with various combinatorial or antibody fragment libraries were conducted on viral targets as well as host receptors to identify HIV-1 inhibitors. Besides these applications, phage display technology has been applied to characterize the enzymatic specificity of the HIV-1 protease. Phage particles also represent valuable alternative carriers displaying various HIV-1 antigens to the immune system and eliciting antiviral responses. This review presents and summarizes the different studies conducted with regard to the nature of phage libraries, target display mode and biopanning procedures.
Keywords: CCR5; CXCR4; HIV-1; HIV-1 inhibitor; HIV-1 vaccine; epitope mapping; gp120; gp41; mimotopes; phage display.
Figures

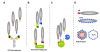

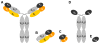
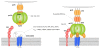
References
-
- Gottlieb M.S., Schroff R., Schanker H.M., Weisman J.D., Fan P.T., Wolf R.A., Saxon A. Pneumocystis carinii pneumonia and mucosal candidiasis in previously healthy homosexual men: Evidence of a new acquired cellular immunodeficiency. N. Engl. J. Med. 1981;305:1425–1431. - PubMed
-
- Barre-Sinoussi F., Chermann J.C., Rey F., Nugeyre M.T., Chamaret S., Gruest J., Dauguet C., Axler-Blin C., Vezinet-Brun F., Rouzioux C., et al. Isolation of a t-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS) Science. 1983;220:868–871. - PubMed
-
- Naeger L.K., Struble K.A., Murray J.S., Birnkrant D.B. Running a tightrope: Regulatory challenges in the development of antiretrovirals. Antiviral Res. 2010;85:232–240. - PubMed
-
- Kilby J.M., Hopkins S., Venetta T.M., DiMassimo B., Cloud G.A., Lee J.Y., Alldredge L., Hunter E., Lambert D., Bolognesi D., et al. Potent suppression of HIV-1 replication in humans by T-20, a peptide inhibitor of gp41-mediated virus entry. Nat. Med. 1998;4:1302–1307. - PubMed
-
- Dorr P., Westby M., Dobbs S., Griffin P., Irvine B., Macartney M., Mori J., Rickett G., Smith-Burchnell C., Napier C., et al. Maraviroc (uk-427,857), a potent, orally bioavailable, and selective small-molecule inhibitor of chemokine receptor CCR5 with broad-spectrum anti-human immunodeficiency virus type 1 activity. Antimicrob. Agents Chemother. 2005;49:4721–4732. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

