Genetic divergence and the genetic architecture of complex traits in chromosome substitution strains of mice
- PMID: 22606935
- PMCID: PMC3406986
- DOI: 10.1186/1471-2156-13-38
Genetic divergence and the genetic architecture of complex traits in chromosome substitution strains of mice
Abstract
Background: The genetic architecture of complex traits strongly influences the consequences of inherited mutations, genetic engineering, environmental and genetic perturbations, and natural and artificial selection. But because most studies are under-powered, the picture of complex traits is often incomplete. Chromosome substitution strains (CSSs) are a unique paradigm for these genome surveys because they enable statistically independent, powerful tests for the phenotypic effects of each chromosome on a uniform inbred genetic background. A previous CSS survey in mice and rats revealed many complex trait genes (QTLs), large phenotypic effects, extensive epistasis, as well as systems properties such as strongly directional phenotypic changes and genetically-determined limits on the range of phenotypic variation. However, the unusually close genetic relation between the CSS progenitor strains in that study raised questions about the impact of genetic divergence: would greater divergence between progenitor strains, with the corresponding changes in gene regulation and protein function, lead to significantly more distinctive phenotypic features, or alternatively would epistasis and systems constraints, which are pervasive in CSSs, limit the range of phenotypic variation regardless of the extent of DNA sequence variation?
Results: We analyzed results for an extensive survey of traits in two new panels of CSSs where the donor strains were derived from inbred strains with more distant origins and discovered a strong similarity in genetic and systems properties among the three CSS panels, regardless of divergence time.
Conclusion: Our results argue that DNA sequence differences between host and donor strains did not substantially affect the architecture of complex traits, and suggest instead that strong epistasis buffered the phenotypic effects of genetic divergence, thereby constraining the range of phenotypic variation.
Figures

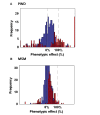
References
-
- Hensen TF. The evolution of genetic architecture. Annu Rev Ecol Evol Syst. 2006;37(1):123–157. doi: 10.1146/annurev.ecolsys.37.091305.110224. - DOI
-
- Saha S, Wu J, Jenkins J, McCarty J, Hayes R, Stelly D. Delineation of interspecific epistasis on fiber quality traits in Gossypium hirsutum by ADAA analysis of intermated G. barbadense chromosome substitution lines. TAG Theor Appl Genet. 2011;122:1351–1361. doi: 10.1007/s00122-011-1536-5. - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

