How genes influence life span: the biodemography of human survival
- PMID: 22607627
- PMCID: PMC3419845
- DOI: 10.1089/rej.2011.1290
How genes influence life span: the biodemography of human survival
Abstract
Background: In genome-wide association studies (GWAS) of human life span, none of the genetic variants has reached the level of genome-wide statistical significance. The roles of such variants in life span regulation remain unclear.
Data and method: A biodemographic analyses was done of genetic regulation of life span using data on low-significance longevity alleles selected in the earlier GWAS of the original Framingham cohort.
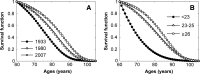
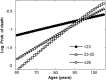
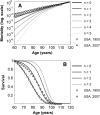
Results: Age-specific survival curves considered as functions of the number of longevity alleles exhibit regularities known in demography as "rectangularization" of survival curves. The presence of such pattern confirms observations from experimental studies that regulation of life span involves genes responsible for stress resistance.
Conclusion: Biodemographic analyses could provide important information about the properties of genes affecting phenotypic traits.
Figures
Similar articles
-
How Genes Modulate Patterns of Aging-Related Changes on the Way to 100: Biodemographic Models and Methods in Genetic Analyses of Longitudinal Data.N Am Actuar J. 2016;20(3):201-232. doi: 10.1080/10920277.2016.1178588. Epub 2016 Jun 22. N Am Actuar J. 2016. PMID: 27773987 Free PMC article.
-
Polygenic effects of common single-nucleotide polymorphisms on life span: when association meets causality.Rejuvenation Res. 2012 Aug;15(4):381-94. doi: 10.1089/rej.2011.1257. Epub 2012 Apr 25. Rejuvenation Res. 2012. PMID: 22533364 Free PMC article.
-
Joint influence of small-effect genetic variants on human longevity.Aging (Albany NY). 2010 Sep;2(9):612-20. doi: 10.18632/aging.100191. Aging (Albany NY). 2010. PMID: 20834067 Free PMC article.
-
The search for longevity and healthy aging genes: insights from epidemiological studies and samples of long-lived individuals.J Gerontol A Biol Sci Med Sci. 2012 May;67(5):470-9. doi: 10.1093/gerona/gls089. Epub 2012 Apr 12. J Gerontol A Biol Sci Med Sci. 2012. PMID: 22499766 Free PMC article. Review.
-
Models to explore genetics of human aging.Adv Exp Med Biol. 2015;847:141-61. doi: 10.1007/978-1-4939-2404-2_7. Adv Exp Med Biol. 2015. PMID: 25916590 Review.
Cited by
-
Mortality increase in late-middle and early-old age: heterogeneity in death processes as a new explanation.Demography. 2013 Oct;50(5):1563-91. doi: 10.1007/s13524-013-0222-4. Demography. 2013. PMID: 23743628 Free PMC article.
-
Joint Analyses of Longitudinal and Time-to-Event Data in Research on Aging: Implications for Predicting Health and Survival.Front Public Health. 2014 Nov 6;2:228. doi: 10.3389/fpubh.2014.00228. eCollection 2014. Front Public Health. 2014. PMID: 25414844 Free PMC article. Review.
-
Roles of interacting stress-related genes in lifespan regulation: insights for translating experimental findings to humans.J Transl Genet Genom. 2021;5(4):357-379. Epub 2021 Oct 19. J Transl Genet Genom. 2021. PMID: 34825130 Free PMC article.
-
Interplay between stress-related genes may influence Alzheimer's disease development: The results of genetic interaction analyses of human data.Mech Ageing Dev. 2021 Jun;196:111477. doi: 10.1016/j.mad.2021.111477. Epub 2021 Mar 30. Mech Ageing Dev. 2021. PMID: 33798591 Free PMC article.
-
Quantification of biological aging in young adults.Proc Natl Acad Sci U S A. 2015 Jul 28;112(30):E4104-10. doi: 10.1073/pnas.1506264112. Epub 2015 Jul 6. Proc Natl Acad Sci U S A. 2015. PMID: 26150497 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

