Degradation of 2,3-dihydroxybenzoate by a novel meta-cleavage pathway
- PMID: 22609919
- PMCID: PMC3416551
- DOI: 10.1128/JB.00430-12
Degradation of 2,3-dihydroxybenzoate by a novel meta-cleavage pathway
Abstract
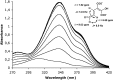
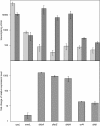
2,3-Dihydroxybenzoate is the precursor in the biosynthesis of several siderophores and an important plant secondary metabolite that, in bacteria, can be degraded via meta-cleavage of the aromatic ring. The dhb cluster of Pseudomonas reinekei MT1 encodes a chimeric meta-cleavage pathway involved in the catabolism of 2,3-dihydroxybenzoate. While the first two enzymes, DhbA and DhbB, are phylogenetically related to those involved in 2,3-dihydroxy-p-cumate degradation, the subsequent steps are catalyzed by enzymes related to those involved in catechol degradation (DhbCDEFGH). Characterization of kinetic properties of DhbA extradiol dioxygenase identified 2,3-dihydroxybenzoate as the preferred substrate. Deletion of the encoding gene impedes growth of P. reinekei MT1 on 2,3-dihydroxybenzoate. DhbA catalyzes 3,4-dioxygenation with 2-hydroxy-3-carboxymuconate as the product, which is then decarboxylated by DhbB to 2-hydroxymuconic semialdehyde. This compound is then subject to dehydrogenation and further degraded to citrate cycle intermediates. Transcriptional analysis revealed genes of the dhB gene cluster to be highly expressed during growth with 2,3-dihydroxybenzoate, whereas a downstream-localized gene encoding 2-hydroxymuconic semialdehyde hydrolase, dispensable for 2,3-dihydroxybenzoate metabolism but crucial for 2,3-dihydroxy-p-cumate degradation, was only marginally expressed. This is the first report describing a gene cluster encoding enzymes for the degradation of 2,3-dihydroxybenzoate.
Figures



References
-
- Arai H, Ohishi T, Chang MY, Kudo T. 2000. Arrangement and regulation of the genes for meta-pathway enzymes required for degradation of phenol in Comamonas testosteroni TA441. Microbiology 146:1707–1715 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

