Endoproteolytic cleavage of TUG protein regulates GLUT4 glucose transporter translocation
- PMID: 22610098
- PMCID: PMC3390669
- DOI: 10.1074/jbc.M112.339457
Endoproteolytic cleavage of TUG protein regulates GLUT4 glucose transporter translocation
Abstract
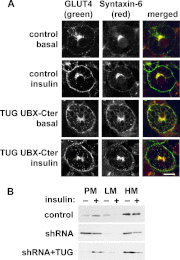
To promote glucose uptake into fat and muscle cells, insulin causes the translocation of GLUT4 glucose transporters from intracellular vesicles to the cell surface. Previous data support a model in which TUG traps GLUT4-containing vesicles and tethers them intracellularly in unstimulated cells and in which insulin mobilizes this pool of vesicles by releasing this tether. Here we show that TUG undergoes site-specific endoproteolytic cleavage, which separates a GLUT4-binding, N-terminal region of TUG from a C-terminal region previously suggested to bind an intracellular anchor. Cleavage is accelerated by insulin stimulation in 3T3-L1 adipocytes and is highly dependent upon adipocyte differentiation. The N-terminal TUG cleavage product has properties of a novel 18-kDa ubiquitin-like modifier, which we call TUGUL. The C-terminal product is observed at the expected size of 42 kDa and also as a 54-kDa form that is released from membranes into the cytosol. In transfected cells, intact TUG links GLUT4 to PIST and also binds Golgin-160 through its C-terminal region. PIST is an effector of TC10α, a GTPase previously shown to transmit an insulin signal required for GLUT4 translocation, and we show using RNAi that TC10α is required for TUG proteolytic processing. Finally, we demonstrate that a cleavage-resistant form of TUG does not support highly insulin-responsive GLUT4 translocation or glucose uptake in 3T3-L1 adipocytes. Together with previous results, these data support a model whereby insulin stimulates TUG cleavage to liberate GLUT4 storage vesicles from the Golgi matrix, which promotes GLUT4 translocation to the cell surface and enhances glucose uptake.
Figures






References
-
- Rubin B. R., Bogan J. S. (2009) Intracellular retention and insulin-stimulated mobilization of GLUT4 glucose transporters. Vitam. Horm. 80, 155–192 - PubMed
-
- Huang S., Czech M. P. (2007) The GLUT4 glucose transporter. Cell Metab. 5, 237–252 - PubMed
-
- Bogan J. S. (2012) Regulation of glucose transporter translocation in health and diabetes. Annu. Rev. Biochem. 81, 507–532 - PubMed
-
- Kandror K. V., Pilch P. F. (2011) The sugar is sIRVed. Sorting Glut4 and its fellow travelers. Traffic 12, 665–671 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

