Evolution of Burkholderia pseudomallei in recurrent melioidosis
- PMID: 22615773
- PMCID: PMC3352902
- DOI: 10.1371/journal.pone.0036507
Evolution of Burkholderia pseudomallei in recurrent melioidosis
Abstract
Burkholderia pseudomallei, the etiologic agent of human melioidosis, is capable of causing severe acute infection with overwhelming septicemia leading to death. A high rate of recurrent disease occurs in adult patients, most often due to recrudescence of the initial infecting strain. Pathogen persistence and evolution during such relapsing infections are not well understood. Bacterial cells present in the primary inoculum and in late infections may differ greatly, as has been observed in chronic disease, or they may be genetically similar. To test these alternative models, we conducted whole-genome comparisons of clonal primary and relapse B. pseudomallei isolates recovered six months to six years apart from four adult Thai patients. We found differences within each of the four pairs, and some, including a 330 Kb deletion, affected substantial portions of the genome. Many of the changes were associated with increased antibiotic resistance. We also found evidence of positive selection for deleterious mutations in a TetR family transcriptional regulator from a set of 107 additional B. pseudomallei strains. As part of the study, we sequenced to base-pair accuracy the genome of B. pseudomallei strain 1026b, the model used for genetic studies of B. pseudomallei pathogenesis and antibiotic resistance. Our findings provide new insights into pathogen evolution during long-term infections and have important implications for the development of intervention strategies to combat recurrent melioidosis.
Conflict of interest statement
Figures
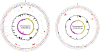


References
-
- Dobrindt U, Zdziarski J, Salvador E, Hacker J. Bacterial genome plasticity and its impact on adaptation during persistent infection. Int J Med Microbiol. 2010;300:363–366. - PubMed
-
- Hogardt M, Heesemann J. Adaptation of Pseudomonas aeruginosa during persistence in the cystic fibrosis lung. Int J Med Microbiol. 2010;300:557–562. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

