Comparative multi-omics systems analysis of Escherichia coli strains B and K-12
- PMID: 22632713
- PMCID: PMC3446290
- DOI: 10.1186/gb-2012-13-5-r37
Comparative multi-omics systems analysis of Escherichia coli strains B and K-12
Abstract
Background: Elucidation of a genotype-phenotype relationship is critical to understand an organism at the whole-system level. Here, we demonstrate that comparative analyses of multi-omics data combined with a computational modeling approach provide a framework for elucidating the phenotypic characteristics of organisms whose genomes are sequenced.
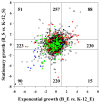
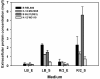
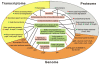
Results: We present a comprehensive analysis of genome-wide measurements incorporating multifaceted holistic data - genome, transcriptome, proteome, and phenome - to determine the differences between Escherichia coli B and K-12 strains. A genome-scale metabolic network of E. coli B was reconstructed and used to identify genetic bases of the phenotypes unique to B compared with K-12 through in silico complementation testing. This systems analysis revealed that E. coli B is well-suited for production of recombinant proteins due to a greater capacity for amino acid biosynthesis, fewer proteases, and lack of flagella. Furthermore, E. coli B has an additional type II secretion system and a different cell wall and outer membrane composition predicted to be more favorable for protein secretion. In contrast, E. coli K-12 showed a higher expression of heat shock genes and was less susceptible to certain stress conditions.
Conclusions: This integrative systems approach provides a high-resolution system-wide view and insights into why two closely related strains of E. coli, B and K-12, manifest distinct phenotypes. Therefore, systematic understanding of cellular physiology and metabolism of the strains is essential not only to determine culture conditions but also to design recombinant hosts.
Figures


Similar articles
-
A comparative analysis of industrial Escherichia coli K-12 and B strains in high-glucose batch cultivations on process-, transcriptome- and proteome level.PLoS One. 2013 Aug 8;8(8):e70516. doi: 10.1371/journal.pone.0070516. eCollection 2013. PLoS One. 2013. PMID: 23950949 Free PMC article.
-
Comparative analysis of envelope proteomes in Escherichia coli B and K-12 strains.J Microbiol Biotechnol. 2012 Apr;22(4):470-8. doi: 10.4014/jmb.1110.10080. J Microbiol Biotechnol. 2012. PMID: 22534293
-
Comprehensive Analysis of Proteomic Differences between Escherichia coli K-12 and B Strains Using Multiplexed Isobaric Tandem Mass Tag (TMT) Labeling.J Microbiol Biotechnol. 2017 Nov 28;27(11):2028-2036. doi: 10.4014/jmb.1708.08024. J Microbiol Biotechnol. 2017. PMID: 28870009
-
Exploring the proteomic characteristics of the Escherichia coli B and K-12 strains in different cellular compartments.J Biosci Bioeng. 2016 Jul;122(1):1-9. doi: 10.1016/j.jbiosc.2015.12.005. Epub 2016 Jan 5. J Biosci Bioeng. 2016. PMID: 26777236 Review.
-
From the sequence to cell modeling: comprehensive functional genomics in Escherichia coli.J Biochem Mol Biol. 2004 Jan 31;37(1):83-92. doi: 10.5483/bmbrep.2004.37.1.083. J Biochem Mol Biol. 2004. PMID: 14761306 Review.
Cited by
-
Interspecific bacterial sensing through airborne signals modulates locomotion and drug resistance.Nat Commun. 2013;4:1809. doi: 10.1038/ncomms2789. Nat Commun. 2013. PMID: 23651997
-
Predicting bacterial growth conditions from mRNA and protein abundances.PLoS One. 2018 Nov 2;13(11):e0206634. doi: 10.1371/journal.pone.0206634. eCollection 2018. PLoS One. 2018. PMID: 30388153 Free PMC article.
-
Combinatorial strategies for improving multiple-stress resistance in industrially relevant Escherichia coli strains.Appl Environ Microbiol. 2014 Oct;80(19):6223-42. doi: 10.1128/AEM.01542-14. Epub 2014 Aug 1. Appl Environ Microbiol. 2014. PMID: 25085490 Free PMC article.
-
Omics strategies for revealing Yersinia pestis virulence.Front Cell Infect Microbiol. 2012 Dec 13;2:157. doi: 10.3389/fcimb.2012.00157. eCollection 2012. Front Cell Infect Microbiol. 2012. PMID: 23248778 Free PMC article. Review.
-
A comprehensive survey of the approaches for pathway analysis using multi-omics data integration.Brief Bioinform. 2022 Nov 19;23(6):bbac435. doi: 10.1093/bib/bbac435. Brief Bioinform. 2022. PMID: 36252928 Free PMC article. Review.
References
-
- Swartz JR. In: Escherichia coli and Salmonella typhimurium: Cellular and Molecular Biology. 2. Neidhardt FC, Curtiss III R, Ingraham JL, Lin ECC, Low KB, Magasanik B, Reznikoff WS, Riley M, Shaechter M, Umbarger HE, editor. Washington, DC: ASM Press; 1996. Escherichia coli recombinant DNA technology. pp. 1693–1712.
-
- Yoon SH, Jeong H, Kwon S-K, Kim JF. Systems Biology and Biotechnology of Escherichia coli. Berlin, Germany: Springer; 2009. Genomics, biological features, and biotechnological applications of Escherichia coli B: "Is B for better?!". pp. 1–17.
-
- Blattner FR, Plunkett G 3rd, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF, Gregor J, Davis NW, Kirkpatrick HA, Goeden MA, Rose DJ, Mau B, Shao Y. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–1474. doi: 10.1126/science.277.5331.1453. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

