Quantification of the affinities and kinetics of protein interactions using silicon nanowire biosensors
- PMID: 22635097
- PMCID: PMC4180882
- DOI: 10.1038/nnano.2012.82
Quantification of the affinities and kinetics of protein interactions using silicon nanowire biosensors
Abstract
Monitoring the binding affinities and kinetics of protein interactions is important in clinical diagnostics and drug development because such information is used to identify new therapeutic candidates. Surface plasmon resonance is at present the standard method used for such analysis, but this is limited by low sensitivity and low-throughput analysis. Here, we show that silicon nanowire field-effect transistors can be used as biosensors to measure protein-ligand binding affinities and kinetics with sensitivities down to femtomolar concentrations. Based on this sensing mechanism, we develop an analytical model to calibrate the sensor response and quantify the molecular binding affinities of two representative protein-ligand binding pairs. The rate constant of the association and dissociation of the protein-ligand pair is determined by monitoring the reaction kinetics, demonstrating that silicon nanowire field-effect transistors can be readily used as high-throughput biosensors to quantify protein interactions.
Figures

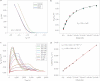
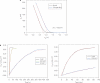

References
-
- Boccaletti S, Latora V, Moreno Y, Chavez M, Hwang DU. Complex networks: structure and dynamics. Phys. Rep. 2006;424:175–308.
-
- Wilson GS, Gifford R. Biosensors for real-time in vivo measurements. Biosens. Bioelectron. 2005;20:2388–2403. - PubMed
-
- Cheng MMC, et al. Nanotechnologies for biomolecular detection and medical diagnostics. Curr. Opin. Chem. Biol. 2006;10:11–19. - PubMed
-
- D'Orazio P. Biosensors in clinical chemistry. Clin. Chim. Acta. 2003;334:41–69. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

