Subunit Interactions and cooperativity in the microtubule-severing AAA ATPase spastin
- PMID: 22637577
- PMCID: PMC3406712
- DOI: 10.1074/jbc.M111.291898
Subunit Interactions and cooperativity in the microtubule-severing AAA ATPase spastin
Abstract
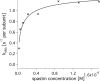
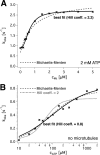
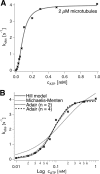
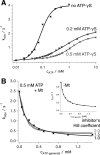
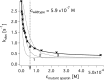
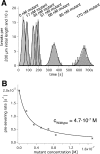
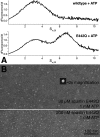
Spastin is a hexameric ring AAA ATPase that severs microtubules. To see whether the ring complex funnels the energy of multiple ATP hydrolysis events to the site of mechanical action, we investigate here the cooperativity of spastin. Several lines of evidence indicate that interactions among two subunits dominate the cooperative behavior: (i) the ATPase activity shows a sigmoidal dependence on the ATP concentration; (ii) ATPγS displays a mixed-inhibition behavior for normal ATP turnover; and (iii) inactive mutant subunits inhibit the activity of spastin in a hyperbolic dependence, characteristic for two interacting species. A quantitative model based on neighbor interactions fits mutant titration experiments well, suggesting that each subunit is mainly influenced by one of its neighbors. These observations are relevant for patients suffering from SPG4-type hereditary spastic paraplegia and explain why single amino acid exchanges lead to a dominant negative phenotype. In severing assays, wild type spastin is even more sensitive toward the presence of inactive mutants than in enzymatic assays, suggesting a weak coupling of ATPase and severing activity.
Figures

References
-
- Errico A., Claudiani P., D'Addio M., Rugarli E. I. (2004) Spastin interacts with the centrosomal protein NA14 and is enriched in the spindle pole, the midbody, and the distal axon. Hum. Mol. Genet. 13, 2121–2132 - PubMed
-
- Svenson I. K., Kloos M. T., Jacon A., Gallione C., Horton A. C., Pericak-Vance M. A., Ehlers M. D., Marchuk D. A. (2005) Subcellular localization of spastin. Implications for the pathogenesis of hereditary spastic paraplegia. Neurogenetics 6, 135–141 - PubMed
-
- Confalonieri F., Duguet M. (1995) A 200-amino acid ATPase module in search of a basic function. BioEssays 17, 639–650 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

