Up for grabs; trashing peroxisomes
- PMID: 22643221
- PMCID: PMC3395098
- DOI: 10.1038/emboj.2012.152
Up for grabs; trashing peroxisomes
Abstract
EMBO J 31 13, 2852–2868 (2012); published online May 29 2012
Together with the proteasome, autophagy is one of the major catabolic pathways of the cell. In response to cellular needs or environmental cues, this transport route targets specific structures for degradation into the mammalian lysosomes or the yeast and plant vacuoles. The mechanisms allowing exclusive autophagic elimination of unwanted structures are currently the object of intensive investigations. The emerging picture is that there is a series of autophagy receptors that determines the specificity of the different selective types of autophagy. How cargo binding and recognition is regulated by these receptors, however, is largely unknown. In their study, Motley et al (2012) have shed light into the molecular principles underlying the turnover of excess peroxisomes in the budding yeast Saccharomyces cerevisiae.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
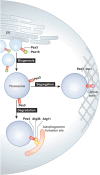
Comment on
-
Pex3-anchored Atg36 tags peroxisomes for degradation in Saccharomyces cerevisiae.EMBO J. 2012 Jun 29;31(13):2852-68. doi: 10.1038/emboj.2012.151. Epub 2012 May 29. EMBO J. 2012. PMID: 22643220 Free PMC article.
References
-
- Bellu AR, Komori M, van der Klei IJ, Kiel JA, Veenhuis M (2001) Peroxisome biogenesis and selective degradation converge at Pex14p. J Biol Chem 276: 44570–44574 - PubMed
-
- Bellu AR, Salomons FA, Kiel JA, Veenhuis M, Van Der Klei IJ (2002) Removal of Pex3p is an important initial stage in selective peroxisome degradation in Hansenula polymorpha. J Biol Chem 277: 42875–42880 - PubMed
-
- Hoepfner D, Schildknegt D, Braakman I, Philippsen P, Tabak HF (2005) Contribution of the endoplasmic reticulum to peroxisome formation. Cell 122: 85–95 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

