Comparison of Atlantic salmon individuals with different outcomes of cardiomyopathy syndrome (CMS)
- PMID: 22646522
- PMCID: PMC3443006
- DOI: 10.1186/1471-2164-13-205
Comparison of Atlantic salmon individuals with different outcomes of cardiomyopathy syndrome (CMS)
Abstract
Background: Cardiomyopathy syndrome (CMS) is a severe disease of Atlantic salmon (Salmo salar L.) associated with significant economic losses in the aquaculture industry. CMS is diagnosed with a severe inflammation and degradation of myocardial tissue caused by a double-stranded RNA virus named piscine myocarditis virus (PMCV), with structural similarities to the Totiviridae family. In the present study we characterized individual host responses and genomic determinants of different disease outcomes.
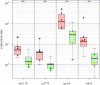
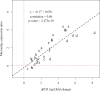
Results: From time course studies of experimentally infected Atlantic salmon post-smolts, fish exhibited different outcomes of infection and disease. High responder (HR) fish were characterized with sustained and increased viral load and pathology in heart tissue. Low responder (LR) fish showed declining viral load from 6-10 weeks post infection (wpi) and absence of pathology. Global gene expression (SIQ2.0 oligonucleotide microarray) in HR and LR hearts during infection was compared, in order to characterize differences in the host response and to identify genes with expression patterns that could explain or predict the different outcomes of disease. Virus-responsive genes involved in early antiviral and innate immune responses were upregulated equally in LR and HR at the first stage (2-4 wpi), reflecting the initial increase in virus replication. Repression of heart muscle development was identified by gene ontology enrichment analyses, indicating the early onset of pathology. By six weeks both responder groups had comparable viral load, while increased pathology was observed in HR fish. This was reflected by induced expression of genes implicated in apoptosis and cell death mechanisms, presumably related to lymphocyte regulation and survival. In contrast, LR fish showed earlier activation of NK cell-mediated cytotoxicity and NOD-like receptor signaling pathways. At the late stage of infection, increased pathology and viral load in HR was accompanied by a broad activation of genes involved in adaptive immunity and particularly T cell responses, probably reflecting the increased infiltration and homing of virus-specific T cells to the infected heart. This was in sharp contrast to LR fish, where recovery and reduced viral load was associated with a significantly reduced transcription of adaptive immunity genes and activation of genes involved in energy metabolism.
Conclusions: In contrast to LR, a stronger and sustained expression of genes involved in adaptive immune responses in heart tissue of HR at the late stage of disease probably reflected the increased lymphocyte infiltration and pathological outcome. In addition to controlled adaptive immunity and activation of genes involved in cardiac energy metabolism in LR at the late stage, recovery of this group could also be related to an earlier activation of NOD-like receptor signaling and NK cell-mediated cytotoxicity pathways.
Figures







References
-
- Brun E, Poppe T, Skrudland A, Jarp J. Cardiomyopathy syndrome in farmed Atlantic salmon Salmo salar: occurrence and direct financial losses for Norwegian aquaculture. Dis Aquat Organ. 2003;56(3):241–247. - PubMed
-
- Amin AB, Trasti J. Endomyocarditis in Atlantic salmon in Norwegian sea farms. Bull Eur Assoc Fish Pathol. 1988;8:70–71.
-
- Ferguson HW, Poppe T, Speare DJ. Cardiomyopathy in farmed Norwegian salmon. Dis Aquat Organ. 1990;8:225–231.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

