Genome rearrangements caused by depletion of essential DNA replication proteins in Saccharomyces cerevisiae
- PMID: 22673806
- PMCID: PMC3430531
- DOI: 10.1534/genetics.112.141051
Genome rearrangements caused by depletion of essential DNA replication proteins in Saccharomyces cerevisiae
Abstract
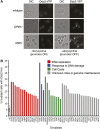
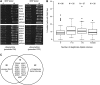
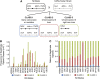
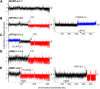
Genetic screens of the collection of ~4500 deletion mutants in Saccharomyces cerevisiae have identified the cohort of nonessential genes that promote maintenance of genome integrity. Here we probe the role of essential genes needed for genome stability. To this end, we screened 217 tetracycline-regulated promoter alleles of essential genes and identified 47 genes whose depletion results in spontaneous DNA damage. We further showed that 92 of these 217 essential genes have a role in suppressing chromosome rearrangements. We identified a core set of 15 genes involved in DNA replication that are critical in preventing both spontaneous DNA damage and genome rearrangements. Mapping, classification, and analysis of rearrangement breakpoints indicated that yeast fragile sites, Ty retrotransposons, tRNA genes, early origins of replication, and replication termination sites are common features at breakpoints when essential replication genes that suppress chromosome rearrangements are downregulated. We propose mechanisms by which depletion of essential replication proteins can lead to double-stranded DNA breaks near these features, which are subsequently repaired by homologous recombination at repeated elements.
Figures

Similar articles
-
Chromosomal translocations in yeast induced by low levels of DNA polymerase a model for chromosome fragile sites.Cell. 2005 Mar 11;120(5):587-98. doi: 10.1016/j.cell.2004.12.039. Cell. 2005. PMID: 15766523
-
Migrating bubble during break-induced replication drives conservative DNA synthesis.Nature. 2013 Oct 17;502(7471):389-92. doi: 10.1038/nature12584. Epub 2013 Sep 11. Nature. 2013. PMID: 24025772 Free PMC article.
-
The F-box protein Dia2 overcomes replication impedance to promote genome stability in Saccharomyces cerevisiae.Genetics. 2006 Dec;174(4):1709-27. doi: 10.1534/genetics.106.057836. Epub 2006 Jun 4. Genetics. 2006. PMID: 16751663 Free PMC article.
-
Recombination between retrotransposons as a source of chromosome rearrangements in the yeast Saccharomyces cerevisiae.DNA Repair (Amst). 2006 Sep 8;5(9-10):1010-20. doi: 10.1016/j.dnarep.2006.05.027. Epub 2006 Jun 23. DNA Repair (Amst). 2006. PMID: 16798113 Review.
-
Prevention of unwanted recombination at damaged replication forks.Curr Genet. 2020 Dec;66(6):1045-1051. doi: 10.1007/s00294-020-01095-7. Epub 2020 Jul 15. Curr Genet. 2020. PMID: 32671464 Free PMC article. Review.
Cited by
-
The Drosophila Mutagen-Sensitivity Gene mus109 Encodes DmDNA2.Genes (Basel). 2022 Feb 7;13(2):312. doi: 10.3390/genes13020312. Genes (Basel). 2022. PMID: 35205357 Free PMC article.
-
Whole Genome Analysis of 132 Clinical Saccharomyces cerevisiae Strains Reveals Extensive Ploidy Variation.G3 (Bethesda). 2016 Aug 9;6(8):2421-34. doi: 10.1534/g3.116.029397. G3 (Bethesda). 2016. PMID: 27317778 Free PMC article.
-
DNA Damage Responses during the Cell Cycle: Insights from Model Organisms and Beyond.Genes (Basel). 2021 Nov 25;12(12):1882. doi: 10.3390/genes12121882. Genes (Basel). 2021. PMID: 34946831 Free PMC article. Review.
-
VID22 counteracts G-quadruplex-induced genome instability.Nucleic Acids Res. 2021 Dec 16;49(22):12785-12804. doi: 10.1093/nar/gkab1156. Nucleic Acids Res. 2021. PMID: 34871443 Free PMC article.
-
Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites.Nat Commun. 2021 Apr 8;12(1):2111. doi: 10.1038/s41467-021-22217-w. Nat Commun. 2021. PMID: 33833229 Free PMC article.
References
-
- Aguilera A., Gomez-Gonzalez B., 2008. Genome instability: a mechanistic view of its causes and consequences. Nat. Rev. Genet. 9: 204–217 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

