Optimized submerged batch fermentation strategy for systems scale studies of metabolic switching in Streptomyces coelicolor A3(2)
- PMID: 22676814
- PMCID: PMC3431225
- DOI: 10.1186/1752-0509-6-59
Optimized submerged batch fermentation strategy for systems scale studies of metabolic switching in Streptomyces coelicolor A3(2)
Abstract
Background: Systems biology approaches to study metabolic switching in Streptomyces coelicolor A3(2) depend on cultivation conditions ensuring high reproducibility and distinct phases of culture growth and secondary metabolite production. In addition, biomass concentrations must be sufficiently high to allow for extensive time-series sampling before occurrence of a given nutrient depletion for transition triggering. The present study describes for the first time the development of a dedicated optimized submerged batch fermentation strategy as the basis for highly time-resolved systems biology studies of metabolic switching in S. coelicolor A3(2).
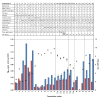
Results: By a step-wise approach, cultivation conditions and two fully defined cultivation media were developed and evaluated using strain M145 of S. coelicolor A3(2), providing a high degree of cultivation reproducibility and enabling reliable studies of the effect of phosphate depletion and L-glutamate depletion on the metabolic transition to antibiotic production phase. Interestingly, both of the two carbon sources provided, D-glucose and L-glutamate, were found to be necessary in order to maintain high growth rates and prevent secondary metabolite production before nutrient depletion. Comparative analysis of batch cultivations with (i) both L-glutamate and D-glucose in excess, (ii) L-glutamate depletion and D-glucose in excess, (iii) L-glutamate as the sole source of carbon and (iv) D-glucose as the sole source of carbon, reveal a complex interplay of the two carbon sources in the bacterium's central carbon metabolism.
Conclusions: The present study presents for the first time a dedicated cultivation strategy fulfilling the requirements for systems biology studies of metabolic switching in S. coelicolor A3(2). Key results from labelling and cultivation experiments on either or both of the two carbon sources provided indicate that in the presence of D-glucose, L-glutamate was the preferred carbon source, while D-glucose alone appeared incapable of maintaining culture growth, likely due to a metabolic bottleneck at the oxidation of pyruvate to acetyl-CoA.
Figures


Similar articles
-
Intracellular glucose 6-phosphate content in Streptomyces coelicolor upon environmental changes in a defined medium.J Biotechnol. 2000 Feb 17;77(2-3):209-18. doi: 10.1016/s0168-1656(99)00215-1. J Biotechnol. 2000. PMID: 10682280
-
Reconstruction of a high-quality metabolic model enables the identification of gene overexpression targets for enhanced antibiotic production in Streptomyces coelicolor A3(2).Biotechnol J. 2014 Sep;9(9):1185-94. doi: 10.1002/biot.201300539. Epub 2014 Apr 23. Biotechnol J. 2014. PMID: 24623710
-
Non-targeted metabolomics unravels a media-dependent prodiginines production pathway in Streptomyces coelicolor A3(2).PLoS One. 2018 Nov 28;13(11):e0207541. doi: 10.1371/journal.pone.0207541. eCollection 2018. PLoS One. 2018. PMID: 30485320 Free PMC article.
-
Multi-level regulation of coelimycin synthesis in Streptomyces coelicolor A3(2).Appl Microbiol Biotechnol. 2019 Aug;103(16):6423-6434. doi: 10.1007/s00253-019-09975-w. Epub 2019 Jun 27. Appl Microbiol Biotechnol. 2019. PMID: 31250060 Free PMC article. Review.
-
Production of specialized metabolites by Streptomyces coelicolor A3(2).Adv Appl Microbiol. 2014;89:217-66. doi: 10.1016/B978-0-12-800259-9.00006-8. Adv Appl Microbiol. 2014. PMID: 25131404 Review.
Cited by
-
Optimized De Novo Eriodictyol Biosynthesis in Streptomyces albidoflavus Using an Expansion of the Golden Standard Toolkit for Its Use in Actinomycetes.Int J Mol Sci. 2023 May 17;24(10):8879. doi: 10.3390/ijms24108879. Int J Mol Sci. 2023. PMID: 37240225 Free PMC article.
-
Mining and fine-tuning sugar uptake system for titer improvement of milbemycins in Streptomyces bingchenggensis.Synth Syst Biotechnol. 2020 Jul 14;5(3):214-221. doi: 10.1016/j.synbio.2020.07.001. eCollection 2020 Sep. Synth Syst Biotechnol. 2020. PMID: 32695892 Free PMC article.
-
Mycelium differentiation and development of Streptomyces coelicolor in lab-scale bioreactors: programmed cell death, differentiation, and lysis are closely linked to undecylprodigiosin and actinorhodin production.Bioresour Technol. 2014 Jan;151:191-8. doi: 10.1016/j.biortech.2013.10.068. Epub 2013 Oct 30. Bioresour Technol. 2014. PMID: 24240146 Free PMC article.
-
Correcting for link loss in causal network inference caused by regulator interference.Bioinformatics. 2014 Oct;30(19):2779-86. doi: 10.1093/bioinformatics/btu388. Epub 2014 Jun 19. Bioinformatics. 2014. PMID: 24947751 Free PMC article.
-
Identification and engineering of 32 membered antifungal macrolactone notonesomycins.Microb Cell Fact. 2020 Mar 19;19(1):71. doi: 10.1186/s12934-020-01328-x. Microb Cell Fact. 2020. PMID: 32192516 Free PMC article.
References
-
- Hopwood DA. Forty years of genetics with Streptomyces: from in vivo through in vitro to in silico. Microbiology. 1999;145:2183–2202. - PubMed
-
- Hopwood DA, Chater KF, Bibb MJ. Genetics of antibiotic production in Streptomyces coelicolor A3(2), a model streptomycete. Biotechnology. 1995;28:65–102. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

