Proofreading of ribonucleotides inserted into DNA by yeast DNA polymerase ɛ
- PMID: 22682724
- PMCID: PMC3407341
- DOI: 10.1016/j.dnarep.2012.05.004
Proofreading of ribonucleotides inserted into DNA by yeast DNA polymerase ɛ
Abstract
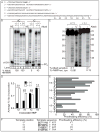
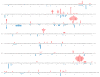
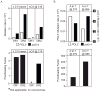
We have investigated the ability of the 3' exonuclease activity of Saccharomyces cerevisiae DNA polymerase ɛ (Pol ɛ) to proofread newly inserted ribonucleotides (rNMPs). During DNA synthesis in vitro, Pol ɛ proofreads ribonucleotides with apparent efficiencies that vary from none at some locations to more than 90% at others, with rA and rU being more efficiently proofread than rC and rG. Previous studies show that failure to repair ribonucleotides in the genome of rnh201Δ strains that lack RNase H2 activity elevates the rate of short deletions in tandem repeat sequences. Here we show that this rate is increased by 2-4-fold in pol2-4 rnh201Δ strains that are also defective in Pol ɛ proofreading. In comparison, defective proofreading in these same strains increases the rate of base substitutions by more than 100-fold. Collectively, the results indicate that although proofreading of an 'incorrect' sugar is less efficient than is proofreading of an incorrect base, Pol ɛ does proofread newly inserted rNMPs to enhance genome stability.
Published by Elsevier B.V.
Conflict of interest statement
None.
Figures

Similar articles
-
Mismatch repair-independent tandem repeat sequence instability resulting from ribonucleotide incorporation by DNA polymerase ε.DNA Repair (Amst). 2011 May 5;10(5):476-82. doi: 10.1016/j.dnarep.2011.02.001. Epub 2011 Mar 16. DNA Repair (Amst). 2011. PMID: 21414850 Free PMC article.
-
Mismatch repair-independent increase in spontaneous mutagenesis in yeast lacking non-essential subunits of DNA polymerase ε.PLoS Genet. 2010 Nov 18;6(11):e1001209. doi: 10.1371/journal.pgen.1001209. PLoS Genet. 2010. PMID: 21124948 Free PMC article.
-
Stimulation of Chromosomal Rearrangements by Ribonucleotides.Genetics. 2015 Nov;201(3):951-61. doi: 10.1534/genetics.115.181149. Epub 2015 Sep 22. Genetics. 2015. PMID: 26400612 Free PMC article.
-
Extrinsic proofreading.DNA Repair (Amst). 2022 Sep;117:103369. doi: 10.1016/j.dnarep.2022.103369. Epub 2022 Jul 4. DNA Repair (Amst). 2022. PMID: 35850061 Free PMC article. Review.
-
Balancing eukaryotic replication asymmetry with replication fidelity.Curr Opin Chem Biol. 2011 Oct;15(5):620-6. doi: 10.1016/j.cbpa.2011.07.025. Epub 2011 Aug 19. Curr Opin Chem Biol. 2011. PMID: 21862387 Free PMC article. Review.
Cited by
-
Human DNA polymerase η has reverse transcriptase activity in cellular environments.J Biol Chem. 2019 Apr 12;294(15):6073-6081. doi: 10.1074/jbc.RA119.007925. Epub 2019 Mar 6. J Biol Chem. 2019. PMID: 30842261 Free PMC article.
-
Eukaryotic genome instability in light of asymmetric DNA replication.Crit Rev Biochem Mol Biol. 2016;51(1):43-52. doi: 10.3109/10409238.2015.1117055. Epub 2015 Dec 20. Crit Rev Biochem Mol Biol. 2016. PMID: 26822554 Free PMC article. Review.
-
Measuring ribonucleotide incorporation into DNA in vitro and in vivo.Methods Mol Biol. 2015;1300:123-39. doi: 10.1007/978-1-4939-2596-4_9. Methods Mol Biol. 2015. PMID: 25916710 Free PMC article.
-
A Critical Balance: dNTPs and the Maintenance of Genome Stability.Genes (Basel). 2017 Jan 31;8(2):57. doi: 10.3390/genes8020057. Genes (Basel). 2017. PMID: 28146119 Free PMC article. Review.
-
Molecular basis for processing of topoisomerase 1-triggered DNA damage by Apn2/APE2.Cell Rep. 2022 Oct 4;41(1):111448. doi: 10.1016/j.celrep.2022.111448. Cell Rep. 2022. PMID: 36198268 Free PMC article.
References
-
- Traut TW. Physiological concentrations of purines and pyrimidines. Mol Cell Biochem. 1994;140:1–22. - PubMed
-
- Chabes A, Georgieva B, Domkin V, Zhao X, Rothstein R, Thelander L. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by relaxed feedback inhibition of ribonucleotide reductase. Cell. 2003;112:391–401. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

