Cloning, production, and functional expression of the bacteriocin enterocin A, produced by Enterococcus faecium T136, by the yeasts Pichia pastoris, Kluyveromyces lactis, Hansenula polymorpha, and Arxula adeninivorans
- PMID: 22685156
- PMCID: PMC3406115
- DOI: 10.1128/AEM.00530-12
Cloning, production, and functional expression of the bacteriocin enterocin A, produced by Enterococcus faecium T136, by the yeasts Pichia pastoris, Kluyveromyces lactis, Hansenula polymorpha, and Arxula adeninivorans
Abstract
The bacteriocin enterocin A (EntA) produced by Enterococcus faecium T136 has been successfully cloned and produced by the yeasts Pichia pastoris X-33EA, Kluyveromyces lactis GG799EA, Hansenula polymorpha KL8-1EA, and Arxula adeninivorans G1212EA. Moreover, P. pastoris X-33EA and K. lactis GG799EA produced EntA in larger amounts and with higher antimicrobial and specific antimicrobial activities than the EntA produced by E. faecium T136.
Figures
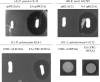
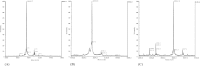
References
-
- Aymerich MT, et al. 2000. Application of enterocins as biopreservatives against Listeria innocua in meat products. J. Food Prot. 63:721–726 - PubMed
-
- Beaulieu L, et al. 2005. Production of pediocin PA-1 in the methylotrophic yeast Pichia pastoris reveals unexpected inhibition of its biological activity due to the presence of collagen-like material. Protein Expr. Purif. 43:111–125 - PubMed
-
- Böer E, Steinborn G, Kunze G, Gellissen G. 2007. Yeast expression platforms. Appl. Microbiol. Biotechnol. 77:513–523 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

