Fast alignment of fragmentation trees
- PMID: 22689771
- PMCID: PMC3371839
- DOI: 10.1093/bioinformatics/bts207
Fast alignment of fragmentation trees
Abstract
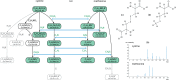
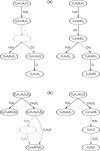
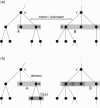
Motivation: Mass spectrometry allows sensitive, automated and high-throughput analysis of small molecules such as metabolites. One major bottleneck in metabolomics is the identification of 'unknown' small molecules not in any database. Recently, fragmentation tree alignments have been introduced for the automated comparison of the fragmentation patterns of small molecules. Fragmentation pattern similarities are strongly correlated with the chemical similarity of the molecules, and allow us to cluster compounds based solely on their fragmentation patterns.
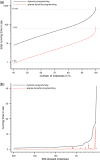
Results: Aligning fragmentation trees is computationally hard. Nevertheless, we present three exact algorithms for the problem: a dynamic programming (DP) algorithm, a sparse variant of the DP, and an Integer Linear Program (ILP). Evaluation of our methods on three different datasets showed that thousands of alignments can be computed in a matter of minutes using DP, even for 'challenging' instances. Running times of the sparse DP were an order of magnitude better than for the classical DP. The ILP was clearly outperformed by both DP approaches. We also found that for both DP algorithms, computing the 1% slowest alignments required as much time as computing the 99% fastest.
Figures
Similar articles
-
Identifying the unknowns by aligning fragmentation trees.Anal Chem. 2012 Apr 3;84(7):3417-26. doi: 10.1021/ac300304u. Epub 2012 Mar 20. Anal Chem. 2012. PMID: 22390817
-
Finding maximum colorful subtrees in practice.J Comput Biol. 2013 Apr;20(4):311-21. doi: 10.1089/cmb.2012.0083. Epub 2013 Mar 19. J Comput Biol. 2013. PMID: 23509858
-
De novo analysis of electron impact mass spectra using fragmentation trees.Anal Chim Acta. 2012 Aug 20;739:67-76. doi: 10.1016/j.aca.2012.06.021. Epub 2012 Jun 27. Anal Chim Acta. 2012. PMID: 22819051
-
Recent advances and prospects of computational methods for metabolite identification: a review with emphasis on machine learning approaches.Brief Bioinform. 2019 Nov 27;20(6):2028-2043. doi: 10.1093/bib/bby066. Brief Bioinform. 2019. PMID: 30099485 Free PMC article. Review.
-
Advances in computational metabolomics and databases deepen the understanding of metabolisms.Curr Opin Biotechnol. 2018 Dec;54:10-17. doi: 10.1016/j.copbio.2018.01.008. Epub 2018 Feb 6. Curr Opin Biotechnol. 2018. PMID: 29413746 Review.
Cited by
-
Using fragmentation trees and mass spectral trees for identifying unknown compounds in metabolomics.Trends Analyt Chem. 2015 Jun 1;69:52-61. doi: 10.1016/j.trac.2015.04.002. Trends Analyt Chem. 2015. PMID: 26213431 Free PMC article.
-
Mass spectrometry tools and workflows for revealing microbial chemistry.Analyst. 2015 Aug 7;140(15):4949-66. doi: 10.1039/c5an00171d. Analyst. 2015. PMID: 25996313 Free PMC article. Review.
-
Computational mass spectrometry for small molecules.J Cheminform. 2013 Mar 1;5(1):12. doi: 10.1186/1758-2946-5-12. J Cheminform. 2013. PMID: 23453222 Free PMC article.
-
Searching molecular structure databases with tandem mass spectra using CSI:FingerID.Proc Natl Acad Sci U S A. 2015 Oct 13;112(41):12580-5. doi: 10.1073/pnas.1509788112. Epub 2015 Sep 21. Proc Natl Acad Sci U S A. 2015. PMID: 26392543 Free PMC article.
References
-
- Arora S., et al. Proof verification and the hardness of approximation problems. J. ACM. 1998;45:501–555.
-
- Backofen R., et al. Sparse RNA folding: time and space efficient algorithms. J. Discrete Algorithms. 2011;9:12–31.
-
- Björklund A., et al. Proceedings of ACM Symposium on Theory of Computing (STOC 2007) New York: ACM Press; 2007. Fourier meets Möbius: fast subset convolution; pp. 67–74.
-
- Böcker S., Rasche F. Towards de novo identification of metabolites by analyzing tandem mass spectra. Bioinformatics. 2008;24:I49–I55. [Proceedings of European Conference on Computational Biology(ECCB 2008)] - PubMed
-
- Canzar S., et al. Proceedings of International Conference on Automata, Languages and Programming (ICALP 2011) Vol. 6755. Berlin: Springer; 2011. On tree-constrained matchings and generalizations; pp. 98–109.

