NPR3 and NPR4 are receptors for the immune signal salicylic acid in plants
- PMID: 22699612
- PMCID: PMC3376392
- DOI: 10.1038/nature11162
NPR3 and NPR4 are receptors for the immune signal salicylic acid in plants
Abstract
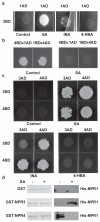
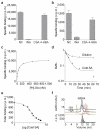
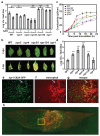
Salicylic acid (SA) is a plant immune signal produced after pathogen challenge to induce systemic acquired resistance. It is the only major plant hormone for which the receptor has not been firmly identified. Systemic acquired resistance in Arabidopsis requires the transcription cofactor nonexpresser of PR genes 1 (NPR1), the degradation of which acts as a molecular switch. Here we show that the NPR1 paralogues NPR3 and NPR4 are SA receptors that bind SA with different affinities. NPR3 and NPR4 function as adaptors of the Cullin 3 ubiquitin E3 ligase to mediate NPR1 degradation in an SA-regulated manner. Accordingly, the Arabidopsis npr3 npr4 double mutant accumulates higher levels of NPR1, and is insensitive to induction of systemic acquired resistance. Moreover, this mutant is defective in pathogen effector-triggered programmed cell death and immunity. Our study reveals the mechanism of SA perception in determining cell death and survival in response to pathogen challenge.
Figures
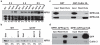
Comment in
-
Plant immunology: A life or death switch.Nature. 2012 Jun 13;486(7402):198-9. doi: 10.1038/486198a. Nature. 2012. PMID: 22699606 No abstract available.
-
How many salicylic acid receptors does a plant cell need?Sci Signal. 2013 Jun 11;6(279):jc3. doi: 10.1126/scisignal.2003944. Sci Signal. 2013. PMID: 23757021
Similar articles
-
Diverse Roles of the Salicylic Acid Receptors NPR1 and NPR3/NPR4 in Plant Immunity.Plant Cell. 2020 Dec;32(12):4002-4016. doi: 10.1105/tpc.20.00499. Epub 2020 Oct 9. Plant Cell. 2020. PMID: 33037144 Free PMC article.
-
UBP12/UBP13-mediated deubiquitination of salicylic acid receptor NPR3 suppresses plant immunity.Mol Plant. 2023 Jan 2;16(1):232-244. doi: 10.1016/j.molp.2022.11.008. Epub 2022 Nov 22. Mol Plant. 2023. PMID: 36415131
-
How many salicylic acid receptors does a plant cell need?Sci Signal. 2013 Jun 11;6(279):jc3. doi: 10.1126/scisignal.2003944. Sci Signal. 2013. PMID: 23757021
-
Perception of the plant immune signal salicylic acid.Curr Opin Plant Biol. 2014 Aug;20:64-8. doi: 10.1016/j.pbi.2014.04.006. Epub 2014 May 20. Curr Opin Plant Biol. 2014. PMID: 24840293 Free PMC article. Review.
-
Progress on the biosynthesis and signal transduction of phytohormone salicylic acid.Yi Chuan. 2020 Sep 20;42(9):858-869. doi: 10.16288/j.yczz.20-173. Yi Chuan. 2020. PMID: 32952120 Review.
Cited by
-
Proteasome-associated ubiquitin ligase relays target plant hormone-specific transcriptional activators.Sci Adv. 2022 Oct 21;8(42):eabn4466. doi: 10.1126/sciadv.abn4466. Epub 2022 Oct 21. Sci Adv. 2022. PMID: 36269824 Free PMC article.
-
Engineering plant immune circuit: walking to the bright future with a novel toolbox.Plant Biotechnol J. 2023 Jan;21(1):17-45. doi: 10.1111/pbi.13916. Epub 2022 Sep 27. Plant Biotechnol J. 2023. PMID: 36036862 Free PMC article. Review.
-
How salicylic acid takes transcriptional control over jasmonic acid signaling.Front Plant Sci. 2015 Mar 25;6:170. doi: 10.3389/fpls.2015.00170. eCollection 2015. Front Plant Sci. 2015. PMID: 25859250 Free PMC article.
-
DELLA proteins and their interacting RING Finger proteins repress gibberellin responses by binding to the promoters of a subset of gibberellin-responsive genes in Arabidopsis.Plant Cell. 2013 Mar;25(3):927-43. doi: 10.1105/tpc.112.108951. Epub 2013 Mar 12. Plant Cell. 2013. PMID: 23482857 Free PMC article.
-
The long-sought-after salicylic acid receptors.Mol Plant. 2012 Sep;5(5):971-3. doi: 10.1093/mp/sss086. Epub 2012 Aug 29. Mol Plant. 2012. PMID: 22933712 Free PMC article. No abstract available.
References
-
- Jones JD, Dangl JL. The plant immune system. Nature. 2006;444:323–329. - PubMed
-
- Lorrain S, Vailleau F, Balague C, Roby D. Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci. 2003;8:263–271. - PubMed
-
- Durrant WE, Dong X. Systemic acquired resistance. Annu Rev Phytopathol. 2004;42:185–209. - PubMed
-
- Chen Z, Silva H, Klessig D. Involvement of reactive oxygen species in the induction of sytemic acquired resistance by salicylic acid in plants. Science. 1994;262:1883–1886. - PubMed
-
- Park SW, Kaimoyo E, Kumar D, Mosher S, Klessig DF. Methyl salicylate is a critical mobile signal for plant systemic acquired resistance. Science. 2007;318:113–116. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

