A histone acetyltransferase regulates active DNA demethylation in Arabidopsis
- PMID: 22700931
- PMCID: PMC3575687
- DOI: 10.1126/science.1219416
A histone acetyltransferase regulates active DNA demethylation in Arabidopsis
Abstract
Active DNA demethylation is an important part of epigenetic regulation in plants and animals. How active DNA demethylation is regulated and its relationship with histone modification patterns are unclear. Here, we report the discovery of IDM1, a regulator of DNA demethylation in Arabidopsis. IDM1 is required for preventing DNA hypermethylation of highly homologous multicopy genes and other repetitive sequences that are normally targeted for active DNA demethylation by Repressor of Silencing 1 and related 5-methylcytosine DNA glycosylases. IDM1 binds methylated DNA at chromatin sites lacking histone H3K4 di- or trimethylation and acetylates H3 to create a chromatin environment permissible for 5-methylcytosine DNA glycosylases to function. Our study reveals how some genes are indicated by multiple epigenetic marks for active DNA demethylation and protection from silencing.
Figures
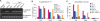


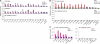
Comment in
-
Molecular biology. All packed up and ready to go.Science. 2012 Jun 15;336(6087):1391-2. doi: 10.1126/science.1224272. Science. 2012. PMID: 22700909 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions

