A nucleotide-gated molecular pore selects sulfotransferase substrates
- PMID: 22703301
- PMCID: PMC3448010
- DOI: 10.1021/bi300631g
A nucleotide-gated molecular pore selects sulfotransferase substrates
Abstract
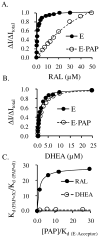
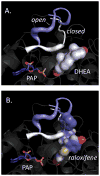
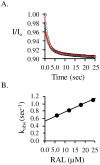
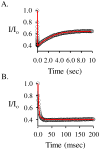
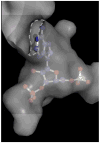
Human SULT2A1 is one of two predominant sulfotransferases in liver and catalyzes transfer of the sulfuryl moiety (-SO(3)) from activated sulfate (PAPS, 3'-phosphoadenosine 5-phosphosulfate) to hundreds of acceptors (metabolites and xenobiotics). Sulfation recodes the biologic activity of acceptors by altering their receptor interactions. The molecular basis on which these enzymes select and sulfonate specific acceptors from complex mixtures of competitors in vivo is a long-standing issue in the SULT field. Raloxifene, a synthetic steroid used in the prevention of osteoporosis, and dehydroepiandrosterone (DHEA), a ubiquitous steroid precusor, are reported to be sulfated efficiently by SULT2A1 in vitro, yet unlike DHEA, raloxifene is not sulfated in vivo. This selectivity was explored in initial rate and equilibrium binding studies that demonstrate pronounced binding antisynergy (21-fold) between PAPS and raloxifene, but not DHEA. Analysis of crystal structures suggests that PAP binding restricts access to the acceptor-binding pocket by restructuring a nine-residue segment of the pocket edge that constricts the active site opening, or "pore", that sieves substrates on the basis of their geometries. In silico docking predicts that raloxifene, which is considerably larger than DHEA, can bind only to the unliganded (open) enzyme, whereas DHEA binds both the open and closed forms. The predictions of these structures with regard to substrate binding are tested using equilibrium and pre-steady-state ligand binding studies, and the results confirm that a nucleotide-driven isomerization controls access to the acceptor-binding pocket and plays an important role in substrate selection by SULT2A1 and possibly other sulfotransferases.
Figures
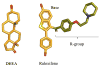

Similar articles
-
Structure, dynamics and selectivity in the sulfotransferase family.Drug Metab Rev. 2013 Nov;45(4):423-30. doi: 10.3109/03602532.2013.835625. Epub 2013 Sep 11. Drug Metab Rev. 2013. PMID: 24025091 Free PMC article. Review.
-
Testing the sulfotransferase molecular pore hypothesis.J Biol Chem. 2013 Mar 22;288(12):8619-8626. doi: 10.1074/jbc.M112.445015. Epub 2013 Jan 28. J Biol Chem. 2013. PMID: 23362278 Free PMC article.
-
Structural rearrangement of SULT2A1: effects on dehydroepiandrosterone and raloxifene sulfation.Horm Mol Biol Clin Investig. 2010;1(2):81-87. doi: 10.1515/HMBCI.2010.012. Horm Mol Biol Clin Investig. 2010. PMID: 21822452 Free PMC article.
-
Paradigms of sulfotransferase catalysis: the mechanism of SULT2A1.J Biol Chem. 2014 Sep 19;289(38):26474-26480. doi: 10.1074/jbc.M114.573501. Epub 2014 Jul 23. J Biol Chem. 2014. PMID: 25056952 Free PMC article.
-
Crystal structure-based studies of cytosolic sulfotransferase.J Biochem Mol Toxicol. 2001;15(2):67-75. doi: 10.1002/jbt.1. J Biochem Mol Toxicol. 2001. PMID: 11284047 Review.
Cited by
-
High accuracy in silico sulfotransferase models.J Biol Chem. 2013 Nov 29;288(48):34494-501. doi: 10.1074/jbc.M113.510974. Epub 2013 Oct 15. J Biol Chem. 2013. PMID: 24129576 Free PMC article.
-
Structure, dynamics and selectivity in the sulfotransferase family.Drug Metab Rev. 2013 Nov;45(4):423-30. doi: 10.3109/03602532.2013.835625. Epub 2013 Sep 11. Drug Metab Rev. 2013. PMID: 24025091 Free PMC article. Review.
-
The multi-protein family of sulfotransferases in plants: composition, occurrence, substrate specificity, and functions.Front Plant Sci. 2014 Oct 16;5:556. doi: 10.3389/fpls.2014.00556. eCollection 2014. Front Plant Sci. 2014. PMID: 25360143 Free PMC article. Review.
-
Sulfotransferase 4A1 Coding Sequence and Protein Structure Are Highly Conserved in Vertebrates.Genes (Basel). 2024 Jul 13;15(7):914. doi: 10.3390/genes15070914. Genes (Basel). 2024. PMID: 39062693 Free PMC article. Review.
-
Binding interactions of hydroxylated polychlorinated biphenyls (OHPCBs) with human hydroxysteroid sulfotransferase hSULT2A1.Chem Biol Interact. 2014 Apr 5;212:56-64. doi: 10.1016/j.cbi.2014.01.018. Epub 2014 Feb 5. Chem Biol Interact. 2014. PMID: 24508592 Free PMC article.
References
-
- Zhang H, Varlamova O, Vargas FM, Falany CN, Leyh TS. Sulfuryl transfer: the catalytic mechanism of human estrogen sulfotransferase. J Biol Chem. 1998;273:10888–10892. - PubMed
-
- Parker CR. Dehydroepiandrosterone and dehydroepiandrosterone sulfate production in the human adrenal during development and aging. Steroids. 1999;64:640–647. - PubMed
-
- Matsubayashi Y, Sakagami Y. Peptide hormones in plants. Annu Rev Plant Biol. 2006;57:649–674. - PubMed
-
- Goldstein DS, Swoboda KJ, Miles JM, Coppack SW, Aneman A, Holmes C, Lamensdorf I, Eisenhofer G. Sources and physiological significance of plasma dopamine sulfate. J Clin Endocrinol Metab. 1999;84:2523–2531. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

