Genomic approaches towards finding cis-regulatory modules in animals
- PMID: 22705667
- PMCID: PMC3541939
- DOI: 10.1038/nrg3242
Genomic approaches towards finding cis-regulatory modules in animals
Abstract
Differential gene expression is the fundamental mechanism underlying animal development and cell differentiation. However, it is a challenge to identify comprehensively and accurately the DNA sequences that are required to regulate gene expression: namely, cis-regulatory modules (CRMs). Three major features, either singly or in combination, are used to predict CRMs: clusters of transcription factor binding site motifs, non-coding DNA that is under evolutionary constraint and biochemical marks associated with CRMs, such as histone modifications and protein occupancy. The validation rates for predictions indicate that identifying diagnostic biochemical marks is the most reliable method, and understanding is enhanced by the analysis of motifs and conservation patterns within those predicted CRMs.
Figures
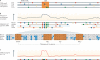
References
-
- Davidson EH, Erwin DH. Gene regulatory networks and the evolution of animal body plans. Science. 2006;311:796–800. - PubMed
-
- King MC, Wilson AC. Evolution at two levels in humans and chimpanzees. Science. 1975;188:107–116. - PubMed
-
- Tuupanen S, et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat Genet. 2009;41:885–890. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

