Glutathione redox potential in the mitochondrial intermembrane space is linked to the cytosol and impacts the Mia40 redox state
- PMID: 22705944
- PMCID: PMC3400016
- DOI: 10.1038/emboj.2012.165
Glutathione redox potential in the mitochondrial intermembrane space is linked to the cytosol and impacts the Mia40 redox state
Abstract
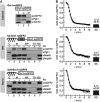
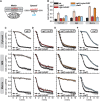
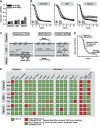
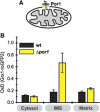
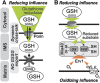
Glutathione is an important mediator and regulator of cellular redox processes. Detailed knowledge of local glutathione redox potential (E(GSH)) dynamics is critical to understand the network of redox processes and their influence on cellular function. Using dynamic oxidant recovery assays together with E(GSH)-specific fluorescent reporters, we investigate the glutathione pools of the cytosol, mitochondrial matrix and intermembrane space (IMS). We demonstrate that the glutathione pools of IMS and cytosol are dynamically interconnected via porins. In contrast, no appreciable communication was observed between the glutathione pools of the IMS and matrix. By modulating redox pathways in the cytosol and IMS, we find that the cytosolic glutathione reductase system is the major determinant of E(GSH) in the IMS, thus explaining a steady-state E(GSH) in the IMS which is similar to the cytosol. Moreover, we show that the local E(GSH) contributes to the partially reduced redox state of the IMS oxidoreductase Mia40 in vivo. Taken together, we provide a comprehensive mechanistic picture of the IMS redox milieu and define the redox influences on Mia40 in living cells.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




Similar articles
-
Kinetic control by limiting glutaredoxin amounts enables thiol oxidation in the reducing mitochondrial intermembrane space.Mol Biol Cell. 2015 Jan 15;26(2):195-204. doi: 10.1091/mbc.E14-10-1422. Epub 2014 Nov 12. Mol Biol Cell. 2015. PMID: 25392302 Free PMC article.
-
Cytosolic Fe-S Cluster Protein Maturation and Iron Regulation Are Independent of the Mitochondrial Erv1/Mia40 Import System.J Biol Chem. 2015 Nov 13;290(46):27829-40. doi: 10.1074/jbc.M115.682179. Epub 2015 Sep 22. J Biol Chem. 2015. PMID: 26396185 Free PMC article.
-
The redox environment in the mitochondrial intermembrane space is maintained separately from the cytosol and matrix.J Biol Chem. 2008 Oct 24;283(43):29126-34. doi: 10.1074/jbc.M803028200. Epub 2008 Aug 15. J Biol Chem. 2008. PMID: 18708636 Free PMC article.
-
Mitochondrial Glutathione: Regulation and Functions.Antioxid Redox Signal. 2017 Nov 20;27(15):1162-1177. doi: 10.1089/ars.2017.7121. Epub 2017 Jun 30. Antioxid Redox Signal. 2017. PMID: 28558477 Review.
-
Structural basis for the disulfide relay system in the mitochondrial intermembrane space.Antioxid Redox Signal. 2010 Nov 1;13(9):1359-73. doi: 10.1089/ars.2010.3099. Antioxid Redox Signal. 2010. PMID: 20136511 Review.
Cited by
-
The mitochondrial disulfide relay system: roles in oxidative protein folding and beyond.Int J Cell Biol. 2013;2013:742923. doi: 10.1155/2013/742923. Epub 2013 Nov 14. Int J Cell Biol. 2013. PMID: 24348563 Free PMC article. Review.
-
Reactive oxygen species and redox compartmentalization.Front Physiol. 2014 Aug 12;5:285. doi: 10.3389/fphys.2014.00285. eCollection 2014. Front Physiol. 2014. PMID: 25161621 Free PMC article. Review.
-
Protein oxidation in the intermembrane space of mitochondria is substrate-specific rather than general.Microb Cell. 2014 Mar 3;1(3):81-93. doi: 10.15698/mic2014.01.130. Microb Cell. 2014. PMID: 28357226 Free PMC article.
-
Protein import and oxidative folding in the mitochondrial intermembrane space of intact mammalian cells.Mol Biol Cell. 2013 Jul;24(14):2160-70. doi: 10.1091/mbc.E12-12-0862. Epub 2013 May 15. Mol Biol Cell. 2013. PMID: 23676665 Free PMC article.
-
Mitochondrial Glutathione in Cellular Redox Homeostasis and Disease Manifestation.Int J Mol Sci. 2024 Jan 21;25(2):1314. doi: 10.3390/ijms25021314. Int J Mol Sci. 2024. PMID: 38279310 Free PMC article. Review.
References
-
- Antos N, Budzinska M, Kmita H (2001) An interplay between the TOM complex and porin isoforms in the yeast Saccharomyces cerevisiae mitochondria. FEBS Lett 500: 12–16 - PubMed
-
- Appenzeller-Herzog C (2011) Glutathione- and non-glutathione-based oxidant control in the endoplasmic reticulum. J Cell Sci 124: 847–855 - PubMed
-
- Banhegyi G, Lusini L, Puskas F, Rossi R, Fulceri R, Braun L, Mile V, di Simplicio P, Mandl J, Benedetti A (1999) Preferential transport of glutathione versus glutathione disulfide in rat liver microsomal vesicles. J Biol Chem 274: 12213–12216 - PubMed
-
- Bien M, Longen S, Wagener N, Chwalla I, Herrmann JM, Riemer J (2010) Mitochondrial disulfide bond formation is driven by intersubunit electron transfer in Erv1 and proofread by glutathione. Mol Cell 37: 516–528 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

