Phylogeography of Camellia taliensis (Theaceae) inferred from chloroplast and nuclear DNA: insights into evolutionary history and conservation
- PMID: 22716114
- PMCID: PMC3495649
- DOI: 10.1186/1471-2148-12-92
Phylogeography of Camellia taliensis (Theaceae) inferred from chloroplast and nuclear DNA: insights into evolutionary history and conservation
Abstract
Background: As one of the most important but seriously endangered wild relatives of the cultivated tea, Camellia taliensis harbors valuable gene resources for tea tree improvement in the future. The knowledge of genetic variation and population structure may provide insights into evolutionary history and germplasm conservation of the species.
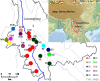
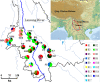
Results: Here, we sampled 21 natural populations from the species' range in China and performed the phylogeography of C. taliensis by using the nuclear PAL gene fragment and chloroplast rpl32-trnL intergenic spacer. Levels of haplotype diversity and nucleotide diversity detected at rpl32-trnL (h = 0.841; π = 0.00314) were almost as high as at PAL (h = 0.836; π = 0.00417). Significant chloroplast DNA population subdivision was detected (GST = 0.988; NST = 0.989), suggesting fairly high genetic differentiation and low levels of recurrent gene flow through seeds among populations. Nested clade phylogeographic analysis of chlorotypes suggests that population genetic structure in C. taliensis has been affected by habitat fragmentation in the past. However, the detection of a moderate nrDNA population subdivision (GST = 0.222; NST = 0.301) provided the evidence of efficient pollen-mediated gene flow among populations and significant phylogeographical structure (NST > GST; P < 0.01). The analysis of PAL haplotypes indicates that phylogeographical pattern of nrDNA haplotypes might be caused by restricted gene flow with isolation by distance, which was also supported by Mantel's test of nrDNA haplotypes (r = 0.234, P < 0.001). We found that chlorotype C1 was fixed in seven populations of Lancang River Region, implying that the Lancang River might have provided a corridor for the long-distance dispersal of the species.
Conclusions: We found that C. taliensis showed fairly high genetic differentiation resulting from restricted gene flow and habitat fragmentation. This phylogeographical study gives us deep insights into population structure of the species and conservation strategies for germplasm sampling and developing in situ conservation of natural populations.
Figures


Similar articles
-
Phylogeography of Chinese cherry (Prunus pseudocerasus Lindl.) inferred from chloroplast and nuclear DNA: insights into evolutionary patterns and demographic history.Plant Biol (Stuttg). 2015 Jul;17(4):787-97. doi: 10.1111/plb.12294. Epub 2015 Jan 9. Plant Biol (Stuttg). 2015. PMID: 25521479
-
Genetic diversity and evolutionary insights of Dali tea (Camellia taliensis) in the Lancang River Basin: Implications for tea breeding and resource conservation.PLoS One. 2025 Jul 31;20(7):e0328658. doi: 10.1371/journal.pone.0328658. eCollection 2025. PLoS One. 2025. PMID: 40743217 Free PMC article.
-
Deciphering the evolutionary imprints of Camellia oleifera Abel.: delineating its distinct phylogeographic structure and demographic history through microsatellite and plastid fragment.BMC Plant Biol. 2025 Apr 25;25(1):526. doi: 10.1186/s12870-025-06570-2. BMC Plant Biol. 2025. PMID: 40275177 Free PMC article.
-
Chloroplast DNA phylogeography of Clintoniaudensis Trautv. & Mey. (Liliaceae) in East Asia.Mol Phylogenet Evol. 2010 May;55(2):721-32. doi: 10.1016/j.ympev.2010.02.010. Epub 2010 Feb 19. Mol Phylogenet Evol. 2010. PMID: 20172032 Review.
-
The diverse applications of cladistic analysis of molecular evolution, with special reference to nested clade analysis.Int J Mol Sci. 2010 Jan 8;11(1):124-39. doi: 10.3390/ijms11010124. Int J Mol Sci. 2010. PMID: 20162005 Free PMC article. Review.
Cited by
-
Biotechnological advances in tea (Camellia sinensis [L.] O. Kuntze): a review.Plant Cell Rep. 2016 Feb;35(2):255-87. doi: 10.1007/s00299-015-1884-8. Epub 2015 Nov 13. Plant Cell Rep. 2016. PMID: 26563347 Review.
-
De novo assembly and transcriptome characterization: novel insights into catechins biosynthesis in Camellia sinensis.BMC Plant Biol. 2014 Oct 15;14:277. doi: 10.1186/s12870-014-0277-4. BMC Plant Biol. 2014. PMID: 25316555 Free PMC article.
-
Complete chloroplast genome of Camellia japonica genome structures, comparative and phylogenetic analysis.PLoS One. 2019 May 9;14(5):e0216645. doi: 10.1371/journal.pone.0216645. eCollection 2019. PLoS One. 2019. PMID: 31071159 Free PMC article.
-
The complete chloroplast genome sequence of rare and endangered Camellia pubipetala Y. Wan & S. Z. Huang (Theaceae) of South China.Mitochondrial DNA B Resour. 2021 Sep 13;6(10):2893-2895. doi: 10.1080/23802359.2021.1915207. eCollection 2021. Mitochondrial DNA B Resour. 2021. PMID: 34532581 Free PMC article.
-
Multiple origins and a narrow genepool characterise the African tea germplasm: concordant patterns revealed by nuclear and plastid DNA markers.Sci Rep. 2017 Jun 22;7(1):4053. doi: 10.1038/s41598-017-04228-0. Sci Rep. 2017. PMID: 28642589 Free PMC article.
References
-
- Hajjar R, Hodgkin T. The use of wild relatives in crop improvement: A survey of developments over the last 20 years. Euphytica. 2007;156:1–13. doi: 10.1007/s10681-007-9363-0. - DOI
-
- Maxted N, Scholten M, Codd R, Ford-Lloyd B. Creation and use of a national inventory of crop wild relatives. Biol Conserv. 2007;140:142–159. doi: 10.1016/j.biocon.2007.08.006. - DOI
-
- Petit RJ, Duminil J, Fineschi S, Hampe A, Salvini D, Vendramin GG. Comparative organization of chloroplast, mitochondrial and nuclear diversity in plant populations. Mol Ecol. 2005;14:689–701. - PubMed
-
- Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA. Phylogeographic studies in plants: problems and prospects. Mol Ecol. 1998;7:465–474. doi: 10.1046/j.1365-294x.1998.00318.x. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

