A two-stage model for lipid modulation of the activity of integral membrane proteins
- PMID: 22723977
- PMCID: PMC3378530
- DOI: 10.1371/journal.pone.0039255
A two-stage model for lipid modulation of the activity of integral membrane proteins
Abstract
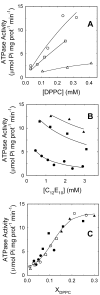
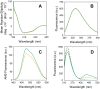
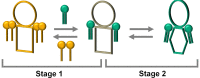
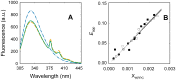
Lipid-protein interactions play an essential role in the regulation of biological function of integral membrane proteins; however, the underlying molecular mechanisms are not fully understood. Here we explore the modulation by phospholipids of the enzymatic activity of the plasma membrane calcium pump reconstituted in detergent-phospholipid mixed micelles of variable composition. The presence of increasing quantities of phospholipids in the micelles produced a cooperative increase in the ATPase activity of the enzyme. This activation effect was reversible and depended on the phospholipid/detergent ratio and not on the total lipid concentration. Enzyme activation was accompanied by a small structural change at the transmembrane domain reported by 1-aniline-8-naphtalenesulfonate fluorescence. In addition, the composition of the amphipilic environment sensed by the protein was evaluated by measuring the relative affinity of the assayed phospholipid for the transmembrane surface of the protein. The obtained results allow us to postulate a two-stage mechanistic model explaining the modulation of protein activity based on the exchange among non-structural amphiphiles at the hydrophobic transmembrane surface, and a lipid-induced conformational change. The model allowed to obtain a cooperativity coefficient reporting on the efficiency of the transduction step between lipid adsorption and catalytic site activation. This model can be easily applied to other phospholipid/detergent mixtures as well to other membrane proteins. The systematic quantitative evaluation of these systems could contribute to gain insight into the structure-activity relationships between proteins and lipids in biological membranes.
Conflict of interest statement
Figures
References
-
- Singer SJ, Nicolson GL. The fluid mosaic model of the structure of cell membranes. Science. 1972;175:720–731. - PubMed
-
- Lee AG. Biological membranes: the importance of molecular detail. Trends in Biochemical Sciences. 2011;36:493–500. - PubMed
-
- Hunte C, Richers S. Lipids and membrane protein structures. Curr Op Struct Biol. 2008;18:406–411. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

