The chromosomal passenger complex controls the function of endosomal sorting complex required for transport-III Snf7 proteins during cytokinesis
- PMID: 22724069
- PMCID: PMC3376741
- DOI: 10.1098/rsob.120070
The chromosomal passenger complex controls the function of endosomal sorting complex required for transport-III Snf7 proteins during cytokinesis
Abstract
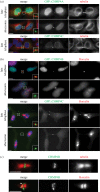
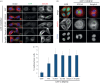
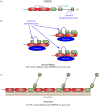
Cytokinesis controls the proper segregation of nuclear and cytoplasmic materials at the end of cell division. The chromosomal passenger complex (CPC) has been proposed to monitor the final separation of the two daughter cells at the end of cytokinesis in order to prevent cell abscission in the presence of DNA at the cleavage site, but the precise molecular basis for this is unclear. Recent studies indicate that abscission could be mediated by the assembly of filaments comprising components of the endosomal sorting complex required for transport-III (ESCRT-III). Here, we show that the CPC subunit Borealin interacts directly with the Snf7 components of ESCRT-III in both Drosophila and human cells. Moreover, we find that the CPC's catalytic subunit, Aurora B kinase, phosphorylates one of the three human Snf7 paralogues-CHMP4C-in its C-terminal tail, a region known to regulate its ability to form polymers and associate with membranes. Phosphorylation at these sites appears essential for CHMP4C function because their mutation leads to cytokinesis defects. We propose that CPC controls abscission timing through inhibition of ESCRT-III Snf7 polymerization and membrane association using two concurrent mechanisms: interaction of its Borealin component with Snf7 proteins and phosphorylation of CHMP4C by Aurora B.
Keywords: Aurora B kinase; Borealin; CHMP4; Shrb; abscission.
Figures




References
-
- Douglas ME, Mishima M. 2010. Still entangled: assembly of the central spindle by multiple microtubule modulators. Semin. Cell Dev. Biol. 21, 899–90810.1016/j.semcdb.2010.08.005 (doi:10.1016/j.semcdb.2010.08.005) - DOI - DOI - PubMed
-
- D'Avino PP, Savoian MS, Glover DM. 2005. Cleavage furrow formation and ingression during animal cytokinesis: a microtubule legacy. J. Cell. Sci. 118, 1549–155810.1242/jcs.02335 (doi:10.1242/jcs.02335) - DOI - DOI - PubMed
-
- Prekeris R, Gould GW. 2008. Breaking up is hard to do: membrane traffic in cytokinesis. J. Cell. Sci. 121, 1569–157610.1242/jcs.018770 (doi:10.1242/jcs.018770) - DOI - DOI - PMC - PubMed
-
- Steigemann P, Gerlich DW. 2009. Cytokinetic abscission: cellular dynamics at the midbody. Trends Cell Biol. 19, 606–61610.1016/j.tcb.2009.07.008 (doi:10.1016/j.tcb.2009.07.008) - DOI - DOI - PubMed
-
- Eggert US, Mitchison TJ, Field CM. 2006. Animal cytokinesis: from parts list to mechanisms. Annu. Rev. Biochem. 75, 543–56610.1146/annurev.biochem.74.082803.133425 (doi:10.1146/annurev.biochem.74.082803.133425) - DOI - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
