Toward almost closed genomes with GapFiller
- PMID: 22731987
- PMCID: PMC3446322
- DOI: 10.1186/gb-2012-13-6-r56
Toward almost closed genomes with GapFiller
Abstract
De novo assembly is a commonly used application of next-generation sequencing experiments. The ultimate goal is to puzzle millions of reads into one complete genome, although draft assemblies usually result in a number of gapped scaffold sequences. In this paper we propose an automated strategy, called GapFiller, to reliably close gaps within scaffolds using paired reads. The method shows good results on both bacterial and eukaryotic datasets, allowing only few errors. As a consequence, the amount of additional wetlab work needed to close a genome is drastically reduced. The software is available at http://www.baseclear.com/bioinformatics-tools/.
Figures
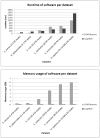

References
-
- Li R, Fan W, Tian G, Zhu H, He L, Cai J, Huang Q, Cai Q, Li B, Bai Y, Zhang Z, Zhang Y, Wang W, Li J, Wei F, Li H, Jian M, Li J, Zhang Z, Nielsen R, Li D, Gu W, Yang Z, Xuan Z, Ryder OA, Leung FC, Zhou Y, Cao J, Sun X, Fu Y. et al. The sequence and de novo assembly of the giant panda genome. Nature. 2010;463:311–317. doi: 10.1038/nature08696. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

